Functions in R Programming
What is a function in R?
A function is a reusable piece of code. It helps you:
- avoid repetition,
- reduce complexity,
- make code easier to test and maintain.
A function can:
- take inputs (arguments),
- execute a body,
- return a value (explicitly with
return()or implicitly as the last expression).
Basic syntax:
Built-in functions (examples)
R ships with many built-in functions. Arguments can be provided by position or by name, and many arguments have defaults.
diff(): compute differences
diff() is often used in time series workflows to compute
lag-1 differences.
set.seed(123)
x <- rnorm(1000)
ts_data <- cumsum(x)
diff_ts <- diff(ts_data)
par(mfrow = c(1, 2))
plot(ts_data, type = "l", main = "Cumulative sum")
plot(diff_ts, type = "l", main = "First differences")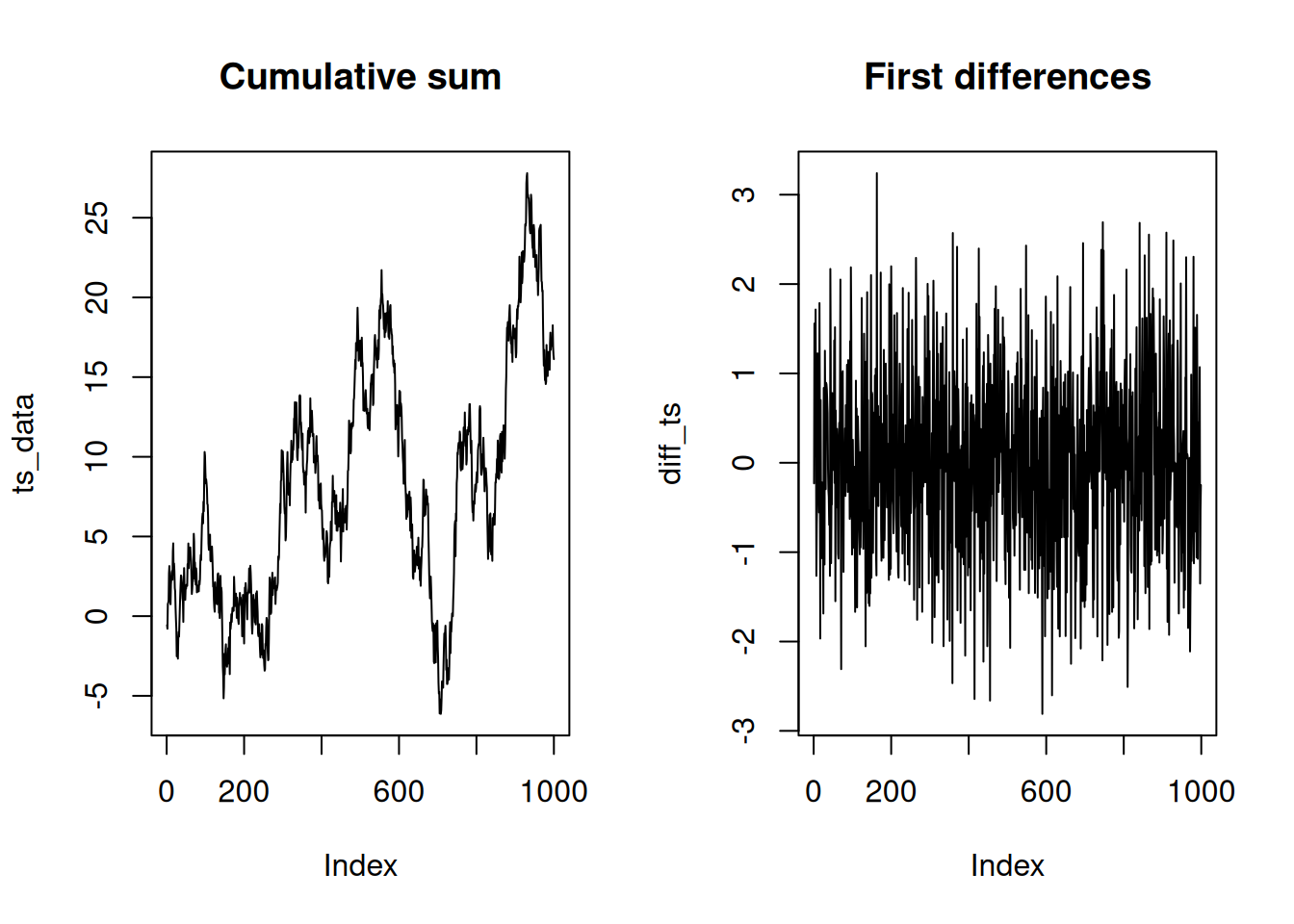
length(): number of elements
length() returns:
- number of elements for vectors,
- number of columns for data frames/matrices.
## [1] 2## [1] 50## [1] 50Math functions
| Function | Description |
|---|---|
| abs(x) | Absolute value |
| log(x, base = b) | Logarithm (natural log if base is omitted) |
| exp(x) | Exponential |
| sqrt(x) | Square root |
| factorial(x) | Factorial |
## [1] 2 0 3## [1] 0.0000000 0.6931472 1.0986123 1.3862944 1.6094379## [1] 2.718282 7.389056 20.085537 54.598150 148.413159## [1] 1.000000 1.414214 1.732051 2.000000 2.236068## [1] 1 2 6 24 120Statistical functions
| Function | Description |
|---|---|
| mean(x) | Mean |
| median(x) | Median |
| var(x) | Variance |
| sd(x) | Standard deviation |
| scale(x) | Z-scores (standardization) |
| quantile(x) | Quantiles |
| summary(x) | Min/1st Qu./Median/Mean/3rd Qu./Max |
## [1] 15.4## [1] 15## [1] 27.95918## [1] 5.287644## [,1]
## [1,] -2.155969
## [2,] -2.155969
## [3,] -1.588609
## [4,] -1.588609
## [5,] -1.399489## 0% 25% 50% 75% 100%
## 4 12 15 19 25## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 4.0 12.0 15.0 15.4 19.0 25.0Writing your own functions
A user-defined function has:
- a name,
- arguments (with optional defaults),
- a body.
One-argument function
## [1] 16## [1] 1 4 9 16 25If you remove a function, check with exists():
## [1] FALSEEnvironments and scoping (lexical scoping)
Functions look for variables in their own local environment first; if not found, they search in the environment where the function was created.
## [1] 15## [1] 10Local variables shadow global ones:
## [1] 105## [1] 10When should you write a function?
If you copy/paste the same logic more than once or twice, it is usually worth writing a function.
Example: normalize to [0, 1]
\[ \text{normalize}(x) = \frac{x - x_{\min}}{x_{\max} - x_{\min}} \]
A robust implementation should handle missing values and constant vectors.
normalize01 <- function(x, na.rm = TRUE) {
rng <- range(x, na.rm = na.rm)
denom <- rng[2] - rng[1]
if (is.na(denom) || denom == 0) {
return(rep(0, length(x)))
}
(x - rng) / denom
}Apply it to multiple columns with modern syntax:
library(tibble)
library(dplyr)
df_example <- tibble(
c1 = rnorm(50, 5, 1.5),
c2 = rnorm(50, 5, 1.5),
c3 = rnorm(50, 5, 1.5)
)
df_example <- df_example |>
mutate(across(starts_with("c"), normalize01, .names = "{.col}_norm"))
df_example |> select(ends_with("_norm")) |> head(5)| c1_norm | c2_norm | c3_norm |
|---|---|---|
| 0.2886787 | 0.6325447 | 0.5629538 |
| -0.7195264 | -0.4891803 | -0.4592776 |
| 0.4703763 | 0.6650164 | 0.2059713 |
| -0.5508433 | -0.6670767 | -0.8834820 |
| 0.0000000 | 0.8255361 | 0.4173301 |
Functions with conditions: train/test split
A more practical split function usually:
- takes a proportion,
- optionally takes a seed for reproducibility,
- returns both train and test sets.
split_data <- function(df, prop = 0.8, seed = NULL) {
stopifnot(is.data.frame(df))
stopifnot(prop > 0 && prop < 1)
if (!is.null(seed)) set.seed(seed)
n <- nrow(df)
n_train <- floor(prop * n)
idx_train <- sample.int(n, size = n_train)
list(
train = df[idx_train, , drop = FALSE],
test = df[-idx_train, , drop = FALSE]
)
}Test it:
data("airquality", package = "datasets")
spl <- split_data(airquality, prop = 0.8, seed = 123)
dim(spl$train)## [1] 122 6## [1] 31 6A work by Gianluca Sottile
gianluca.sottile@unipa.it