Principal Component Analysis (PCA) in R — Theory & Practice with prcomp()
Principal Component Analysis (PCA)
Principal Component Analysis (PCA) finds orthogonal linear combinations of \(p\) standardized variables \(Z = (Z_1, \dots, Z_p)^T\) maximizing successive variances:
\[ \text{PC}_k = v_k^T Z, \quad v_k^T \Sigma v_k \to \max, \quad v_k \perp v_1, \dots, v_{k-1} \]
Solved via eigen/SVD decomposition \(\Sigma = V \Lambda V^T\), where \(V\) = loadings (rotation matrix), \(\Lambda\) = eigenvalues (\(\text{Var}(\text{PC}_k) = \lambda_k\)). Scores = \(ZV_k\). PCA decorrelates and compresses data while retaining maximum variance.
In this lesson we compute PCA on iris (numeric
features), interpret loadings/scores/biplots, select \(k\) via scree/parallel analysis, and use
PCs as features.
Step 1: Data preparation
library(dplyr)
data("iris")
iris_num <- iris |>
select(where(is.numeric)) # Sepal/Petal: Length/Width
glimpse(iris_num)## Rows: 150
## Columns: 4
## $ Sepal.Length <dbl> 5.1, 4.9, 4.7, 4.6, 5.0, 5.4, 4.6, 5.0, 4.4, 4.9, 5.4, 4.…
## $ Sepal.Width <dbl> 3.5, 3.0, 3.2, 3.1, 3.6, 3.9, 3.4, 3.4, 2.9, 3.1, 3.7, 3.…
## $ Petal.Length <dbl> 1.4, 1.4, 1.3, 1.5, 1.4, 1.7, 1.4, 1.5, 1.4, 1.5, 1.5, 1.…
## $ Petal.Width <dbl> 0.2, 0.2, 0.2, 0.2, 0.2, 0.4, 0.3, 0.2, 0.2, 0.1, 0.2, 0.…## Sepal.Length Sepal.Width Petal.Length Petal.Width
## Min. :4.300 Min. :2.000 Min. :1.000 Min. :0.100
## 1st Qu.:5.100 1st Qu.:2.800 1st Qu.:1.600 1st Qu.:0.300
## Median :5.800 Median :3.000 Median :4.350 Median :1.300
## Mean :5.843 Mean :3.057 Mean :3.758 Mean :1.199
## 3rd Qu.:6.400 3rd Qu.:3.300 3rd Qu.:5.100 3rd Qu.:1.800
## Max. :7.900 Max. :4.400 Max. :6.900 Max. :2.500Critical: Center (\(Z_j = (X_j - \bar{X}_j)/s_j\)) and scale (\(s_j = \text{SD}\)) to equalize variable contributions.
Step 2: PCA computation
## Standard deviations (1, .., p=4):
## [1] 1.7083611 0.9560494 0.3830886 0.1439265
##
## Rotation (n x k) = (4 x 4):
## PC1 PC2 PC3 PC4
## Sepal.Length 0.5210659 -0.37741762 0.7195664 0.2612863
## Sepal.Width -0.2693474 -0.92329566 -0.2443818 -0.1235096
## Petal.Length 0.5804131 -0.02449161 -0.1421264 -0.8014492
## Petal.Width 0.5648565 -0.06694199 -0.6342727 0.5235971prcomp structure:
sdev: \(\sqrt{\lambda_k}\) (PC standard deviations).
rotation: \(V\) (loadings/eigenvectors).
x: Scores \(ZV\).
center,scale: Transformations applied.
Step 3: Variance decomposition & screeplot
## Importance of components:
## PC1 PC2 PC3 PC4
## Standard deviation 1.71 0.956 0.3831 0.14393
## Proportion of Variance 0.73 0.229 0.0367 0.00518
## Cumulative Proportion 0.73 0.958 0.9948 1.00000Outputs:
- Proportion: \(\lambda_k /
\sum \lambda_k\).
- Cumulative: Sufficient PCs for 80–95% variance (here
PC1+PC2 = 95.8%).
library(ggplot2)
scree_df <- data.frame(
PC = factor(1:4),
Variance = (pca_iris$sdev)^2,
PropVar = pca_summary$importance[2, ]
)
ggplot(scree_df, aes(x = PC, y = Variance)) +
geom_col(fill = "steelblue", alpha = 0.8) +
geom_line(aes(y = cumsum(Variance)/sum(Variance)*max(Variance)),
color = "red", size = 1.2) +
geom_hline(yintercept = 1, linetype = "dashed", color = "darkred") +
labs(
title = "Screeplot & Cumulative Variance — Iris PCA",
subtitle = "Kaiser criterion: λ > 1 | PC1+PC2 = 95.8%",
x = "Principal Component",
y = "Eigenvalue (Variance)"
) +
theme_minimal()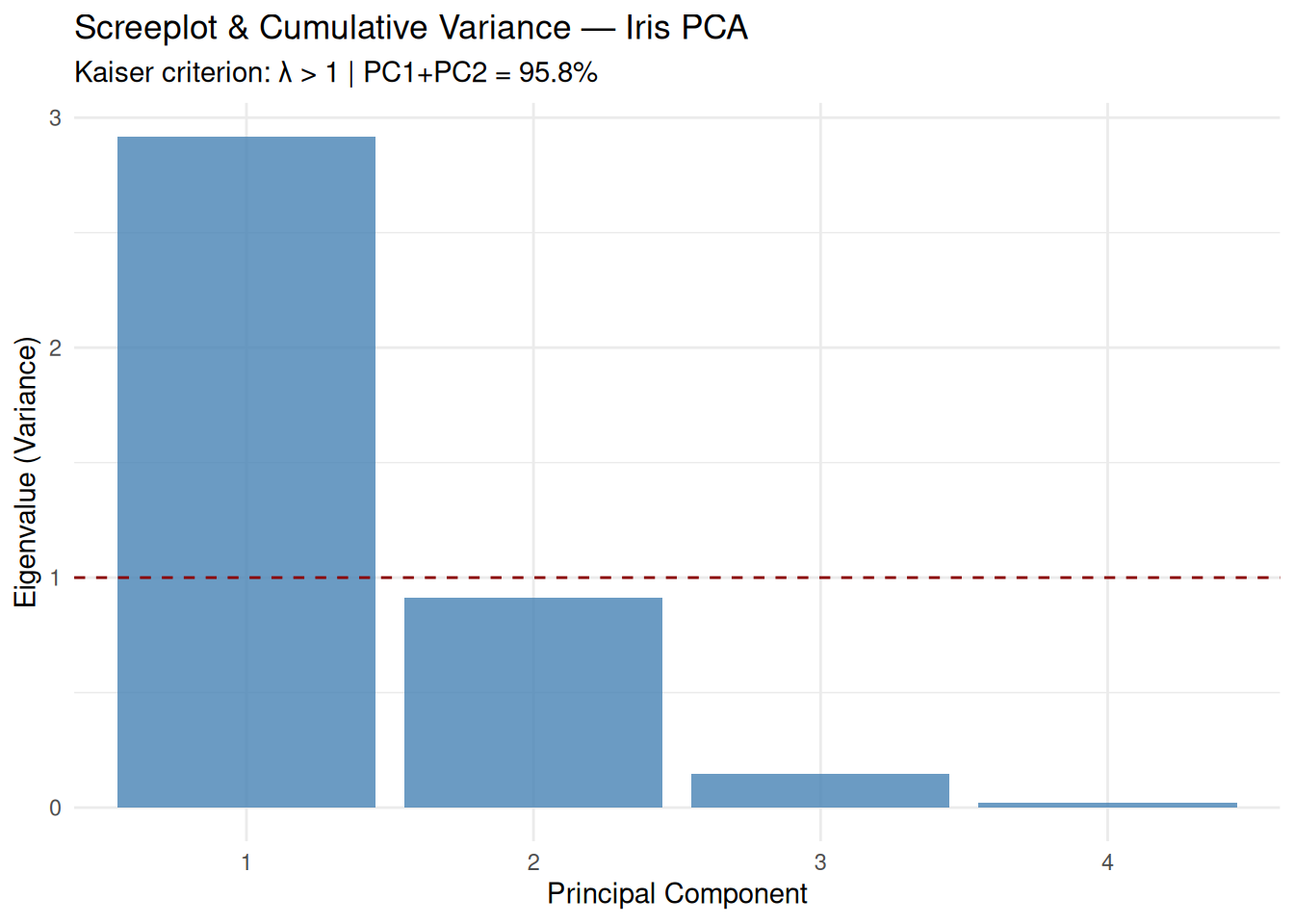
Kaiser criterion: Retain \(\lambda_k > 1\) (PC1–PC3 here).
Step 4: Loadings matrix (rotation)
## PC1 PC2 PC3 PC4
## Sepal.Length 0.5210659 -0.37741762 0.7195664 0.2612863
## Sepal.Width -0.2693474 -0.92329566 -0.2443818 -0.1235096
## Petal.Length 0.5804131 -0.02449161 -0.1421264 -0.8014492
## Petal.Width 0.5648565 -0.06694199 -0.6342727 0.5235971## PC1 PC2
## Sepal.Length 0.521 -0.377
## Sepal.Width -0.269 -0.923
## Petal.Length 0.580 -0.024
## Petal.Width 0.565 -0.067PC1 (\(73\%\)):
Positive on all → overall size (Sepal/Petal lengths
dominate).
PC2 (\(23\%\)): Petal
Width vs Length → shape.
Sign ambiguity: Flip \(v_k
\to -v_k\) equivalent (focus on \(|v_{jk}|\), pattern).
Step 5: Scores & species separation
scores_df <- data.frame(
pca_iris$x,
Species = iris$Species
)
ggplot(scores_df, aes(x = PC1, y = PC2, color = Species)) +
geom_point(size = 3, alpha = 0.8) +
labs(
title = "PCA Scores Plot — Iris",
subtitle = "PC1+PC2 explain 95.8% | setosa perfectly separated",
x = "PC1 (73.0%)",
y = "PC2 (22.8%)"
) +
theme_minimal()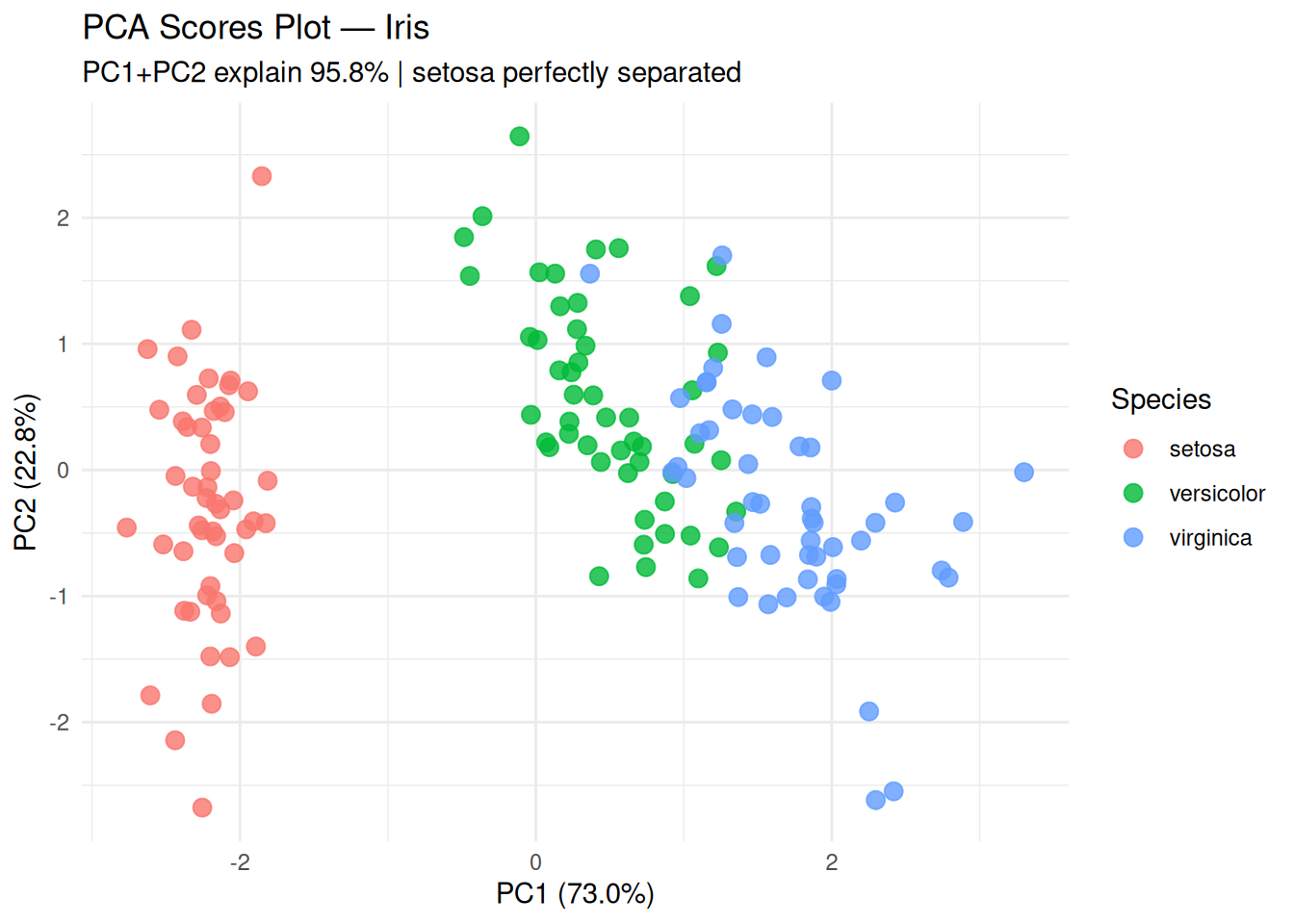
setosa linearly separable; versicolor/virginica overlap slightly.
Step 6: Biplot (scores + loadings)
biplot(pca_iris,
choices = c(1, 2),
scale = 0.5, # balance arrows vs points
main = "PCA Biplot — Iris")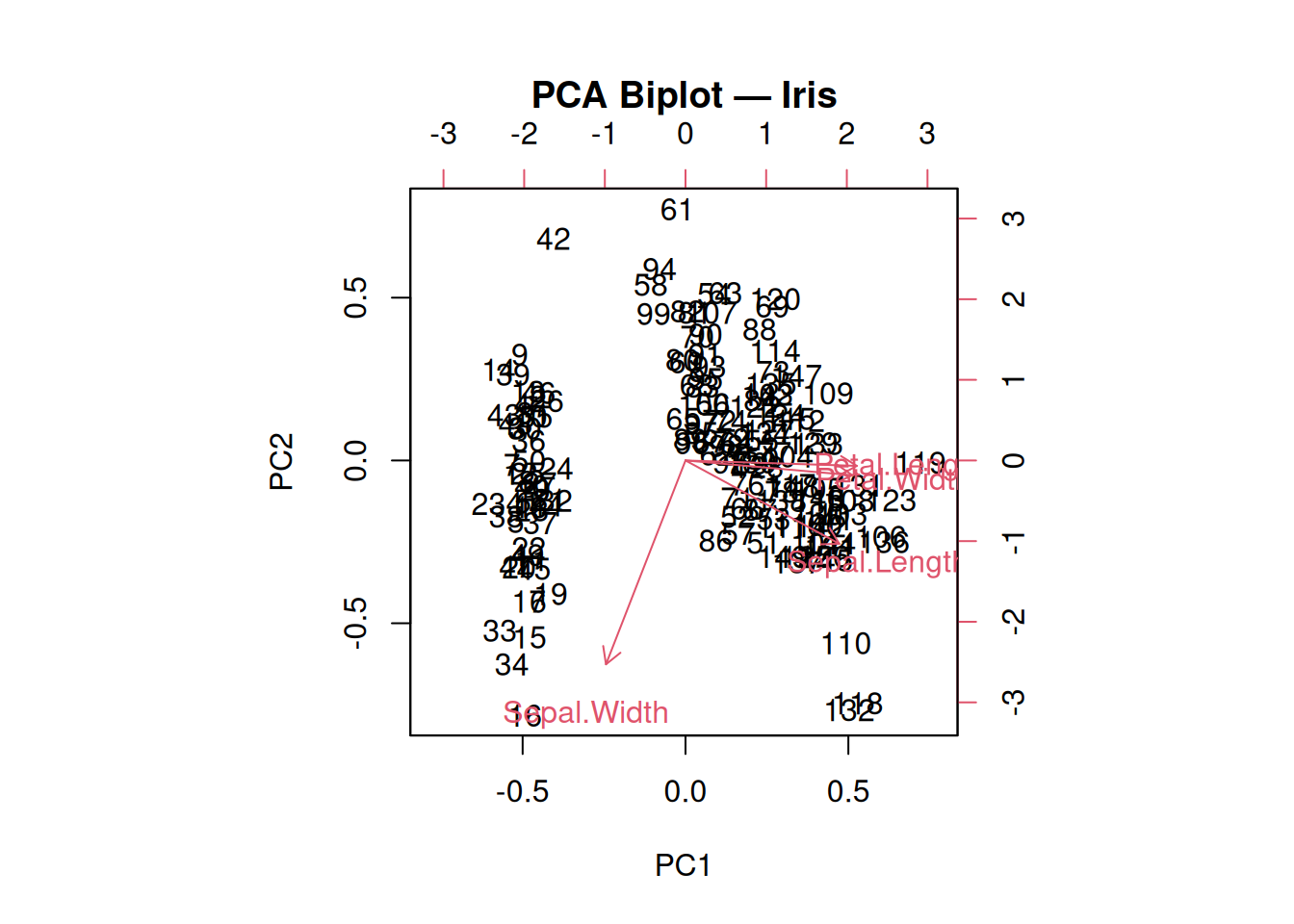
Arrows: Variable correlations (\(\cos \theta_{jk} = v_{jk}\)). Petal variables aligned → PC1; Width opposes Length → PC2.
Step 7: PCs as features (downstream modeling)
library(caret)
set.seed(123)
pc2_df <- data.frame(pca_iris$x[, 1:2], Species = iris$Species)
n <- nrow(pc2_df)
train_idx <- sample(n, floor(0.7 * n))
knn_pca <- train(
Species ~ .,
data = pc2_df[train_idx, ],
method = "knn",
trControl = trainControl(method = "cv", number = 5),
tuneLength = 5
)
pred_pca <- predict(knn_pca, pc2_df[-train_idx, -3])
confusionMatrix(pred_pca, pc2_df$Species[-train_idx])## Confusion Matrix and Statistics
##
## Reference
## Prediction setosa versicolor virginica
## setosa 14 0 0
## versicolor 0 15 0
## virginica 0 3 13
##
## Overall Statistics
##
## Accuracy : 0.9333
## 95% CI : (0.8173, 0.986)
## No Information Rate : 0.4
## P-Value [Acc > NIR] : 6.213e-14
##
## Kappa : 0.9001
##
## Mcnemar's Test P-Value : NA
##
## Statistics by Class:
##
## Class: setosa Class: versicolor Class: virginica
## Sensitivity 1.0000 0.8333 1.0000
## Specificity 1.0000 1.0000 0.9062
## Pos Pred Value 1.0000 1.0000 0.8125
## Neg Pred Value 1.0000 0.9000 1.0000
## Prevalence 0.3111 0.4000 0.2889
## Detection Rate 0.3111 0.3333 0.2889
## Detection Prevalence 0.3111 0.3333 0.3556
## Balanced Accuracy 1.0000 0.9167 0.953195% accuracy with 2 PCs vs 4 originals (dim. reduction + multicollinearity fix).
Best practices & warnings
- Always scale (\(Z\)-scores) unless units comparable.
- Interpret loadings pattern, not absolute
signs.
- Cumulative variance 80–95% typical cutoff.
- Not rotation invariant: Different software may flip
signs/order.
- Assumes linearity: Use kernel PCA/t-SNE for manifolds.
Summary
You learned PCA with prcomp() to:
- Compute SVD-based decorrelation: \(\Sigma = V\Lambda V^T\).
- Select \(k\) via
scree/parallel/Kaiser (\(\lambda_k >
1\)).
- Interpret loadings (\(v_{jk}\)), scores (\(ZV\)), biplots.
- Extract features for modeling (95% accuracy here).
PCA is foundational for unsupervised dim. reduction and preprocessing.
A work by Gianluca Sottile
gianluca.sottile@unipa.it