DBSCAN in R — eps, minPts, Noise Handling, and Non-spherical Clusters
DBSCAN (Density-Based Clustering)
DBSCAN groups points based on density rather than distance to centroids. Key concepts:
- \(\varepsilon\)-neighborhood: \(N_\varepsilon(x) = \{y: \|x-y\| \le \varepsilon\}\).
- Core point: \(|N_\varepsilon(x)| \ge \text{minPts}\).
- Border point: not core but density-reachable from a core point.
- Noise: points not density-reachable from any core point.
DBSCAN can detect non-spherical clusters and identifies outliers/noise explicitly.
In this lesson
- We preprocess business data as in previous lesson.
- We choose
epsusing kNN distance plot. - We run DBSCAN and interpret clusters vs noise.
- We compare DBSCAN with K-means on the non-noise subset.
Step 1: Import + preprocessing
library(readr)
library(dplyr)
local_path <- "raw_data/wholesale_customers.csv"
if (!file.exists(local_path)) {
dir.create("raw_data", showWarnings = FALSE)
download.file(url_path, local_path, mode = "wb")
}
df_raw <- read_csv(local_path, show_col_types = FALSE)
spend_vars <- c("Fresh", "Milk", "Grocery", "Frozen", "Detergents_Paper", "Delicassen")
X <- df_raw |>
mutate(across(all_of(spend_vars), ~ log1p(.x))) |>
select(all_of(spend_vars)) |>
mutate(across(everything(), scale)) |>
as.data.frame()Step 2: Parameter selection (minPts and eps)
A common heuristic sets \(\text{minPts} \approx 2p\) where \(p\) is the number of features.
## [1] 12To choose eps, inspect the kNN distance plot: pick
eps near the “knee”.
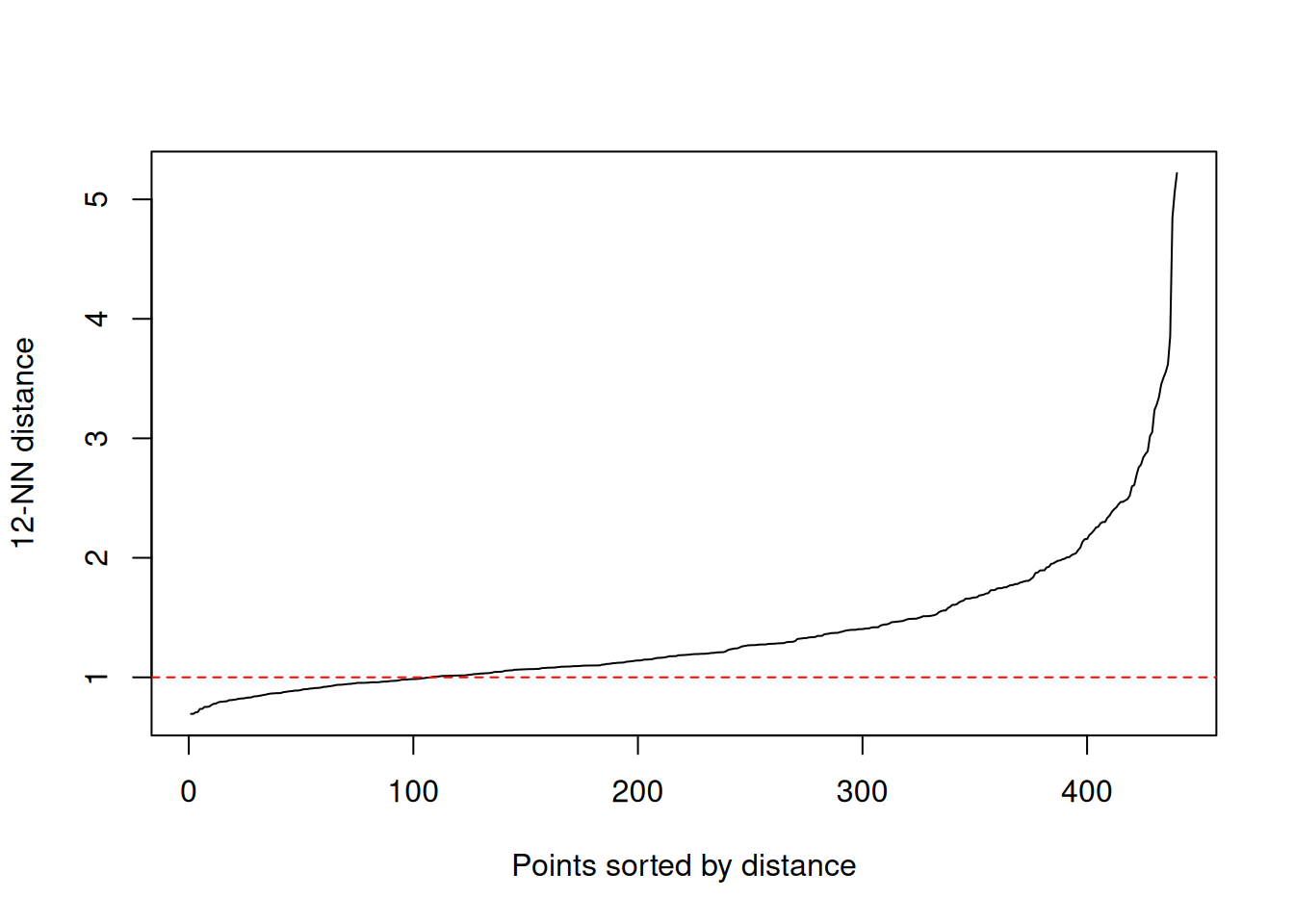
Replace the horizontal line with your candidate value for
eps after inspecting the knee.
Step 3: Fit DBSCAN
## DBSCAN clustering for 440 objects.
## Parameters: eps = 1, minPts = 12
## Using euclidean distances and borderpoints = TRUE
## The clustering contains 1 cluster(s) and 182 noise points.
##
## 0 1
## 182 258
##
## Available fields: cluster, eps, minPts, metric, borderPoints##
## 0 1
## 182 258Interpretation: - If most points are noise, eps may be
too small or minPts too large. - If almost no noise and very few
clusters, eps may be too large (merging structures).
Step 4: Visualization (PCA projection)
library(ggplot2)
pc <- prcomp(X)
plot_df <- data.frame(
PC1 = pc$x[, 1],
PC2 = pc$x[, 2],
cluster = factor(db$cluster)
)
ggplot(plot_df, aes(PC1, PC2, color = cluster)) +
geom_point(alpha = 0.8, size = 2) +
theme_minimal() +
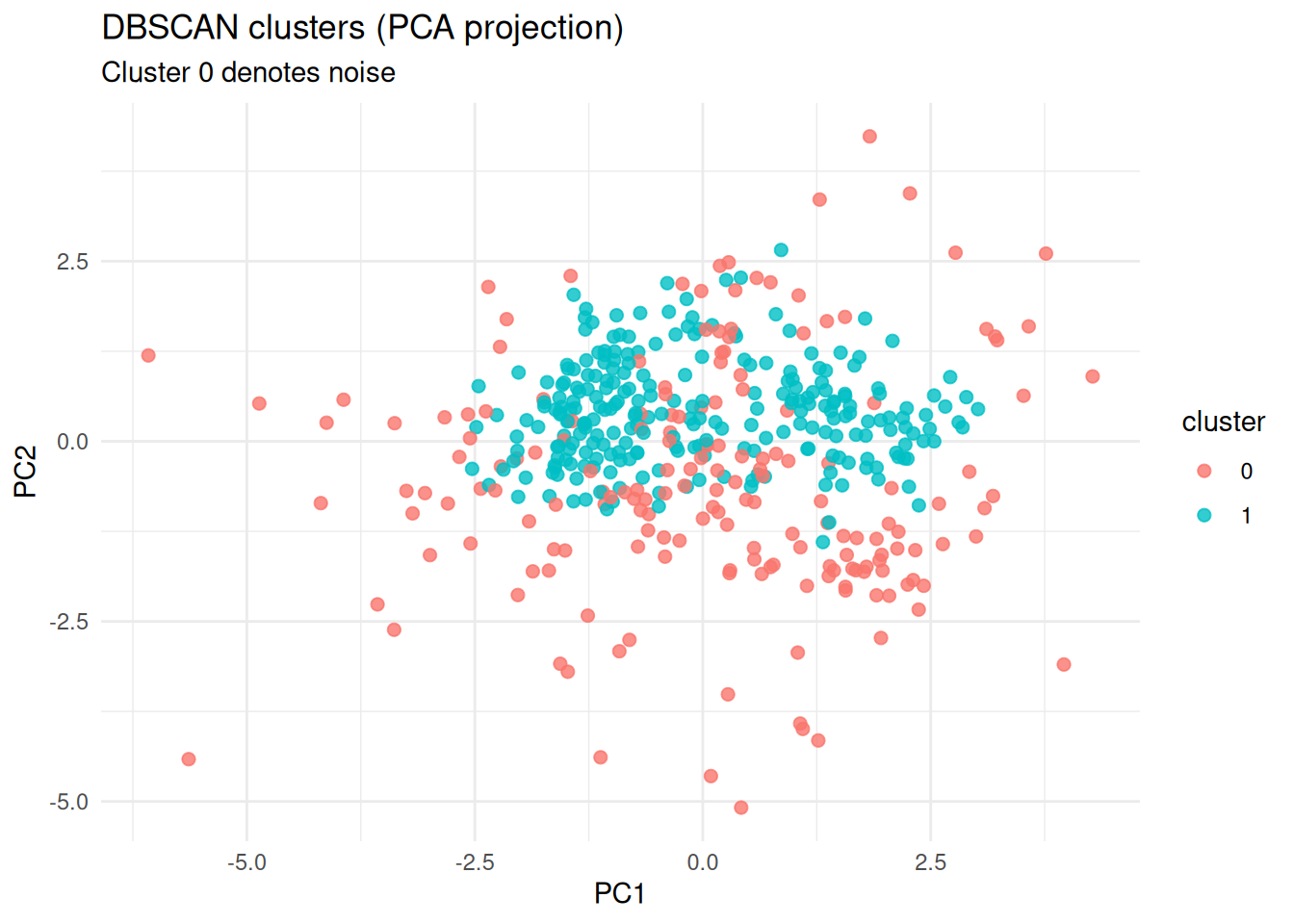
labs(
title = "DBSCAN clusters (PCA projection)",
subtitle = "Cluster 0 denotes noise",
x = "PC1", y = "PC2"
)
Step 5: Internal validation (silhouette on non-noise points)
Silhouette is not directly defined for noise. A practical approach is to compute silhouette on the subset with cluster label \(\ge 1\).
Step 6: Comparison with K-means (on non-noise subset)
library(mclust)
k_db <- length(unique(cl_nonoise))
set.seed(123)
km <- kmeans(X_nonoise, centers = k_db, nstart = 50)
ari <- adjustedRandIndex(cl_nonoise, km$cluster)
ari## [1] 1Interpretation:
- Agreement is not expected to be perfect, because DBSCAN optimizes density connectivity, not SSE.
- Differences often highlight non-spherical structure or outliers.
Step 7: Practical reporting
For a business segmentation report:
- Report number of clusters found (excluding noise) and proportion of noise.
- Provide cluster profiles (feature means in original units, not only z-scores).
- Explain that DBSCAN can isolate atypical customers as noise/outliers, which may be a feature rather than a drawback.
Summary
- DBSCAN is density-based and can recover non-spherical clusters.
minPtsandepsare critical; the kNN distance plot is a practical tool for selectingeps.- Noise labeling provides a principled way to treat outliers.
- Compare with K-means on the non-noise subset to understand structural differences.
A work by Gianluca Sottile
gianluca.sottile@unipa.it