Model-based Clustering in R — Gaussian Mixtures, Soft Assignments, and Component Selection
Model-based clustering (Gaussian Mixture Models)
Model-based clustering assumes data arise from a mixture distribution: \(p(x) = \sum_{g=1}^G \pi_g \, \phi(x \mid \mu_g, \Sigma_g)\), where \(\pi_g \ge 0\) and \(\sum_g \pi_g = 1\).
Key properties:
- Soft assignments via posterior probabilities: \(\tau_{ig} = P(z_i=g \mid x_i)\).
- Clusters can be elliptical (through \(\Sigma_g\)), unlike K-means which is inherently spherical in Euclidean space.
- The number of components \(G\) can be selected using information criteria (commonly BIC).
In this lesson:
- Fit a GMM with automatic model selection.
- Inspect soft assignments (uncertainty).
- Compare with K-means.
Step 1: Import + preprocessing
library(readr)
library(dplyr)
local_path <- "raw_data/wholesale_customers.csv"
df_raw <- read_csv(local_path, show_col_types = FALSE)
spend_vars <- c("Fresh", "Milk", "Grocery", "Frozen", "Detergents_Paper", "Delicassen")
X <- df_raw |>
mutate(across(all_of(spend_vars), ~ log1p(.x))) |>
select(all_of(spend_vars)) |>
mutate(across(everything(), scale)) |>
as.matrix()Step 2: Fit model-based clustering and select G by BIC
# install.packages("mclust") # if needed
library(mclust)
set.seed(123)
fit <- Mclust(X, G = 1:12, verbose = FALSE)
fit## 'Mclust' model object: (VVE,5)
##
## Available components:
## [1] "call" "data" "modelName" "n"
## [5] "d" "G" "BIC" "loglik"
## [9] "df" "bic" "icl" "hypvol"
## [13] "parameters" "z" "classification" "uncertainty"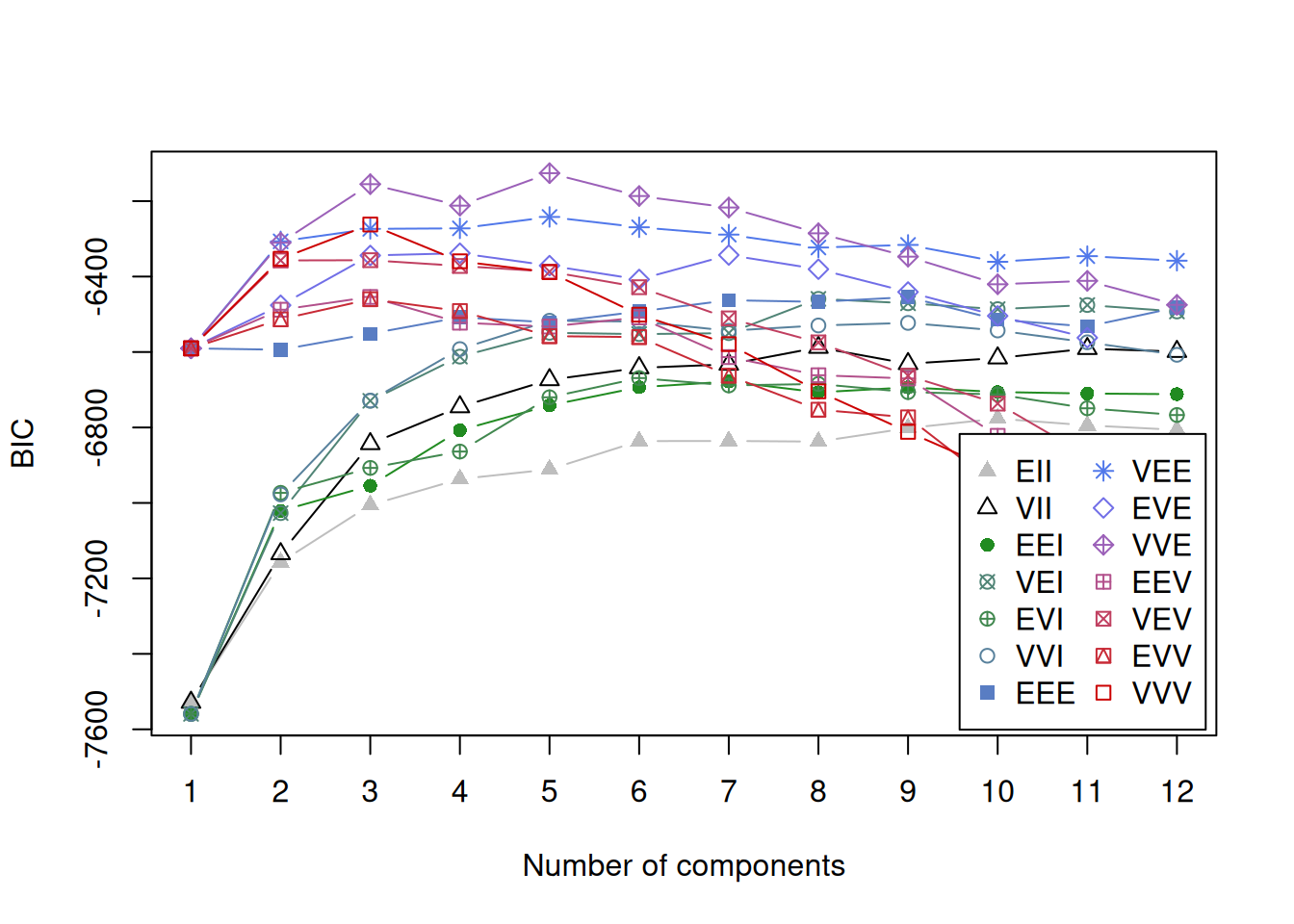
Commentary:
- If the BIC curve has a clear maximum, the selected \(G\) is usually stable.
- If BIC is flat across many \(G\), clustering structure may be weak or sensitive to preprocessing.
Step 3: Hard vs soft clustering outputs
cl <- fit$classification # hard labels
prob <- fit$z # posterior probabilities (n x G)
uncert <- 1 - apply(prob, 1, max) # simple uncertainty measure
table(cl)## cl
## 1 2 3 4 5
## 51 101 155 81 52## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 0.00000 0.01345 0.04573 0.11402 0.16097 0.60119High uncertainty points indicate observations not clearly explained by any single component.
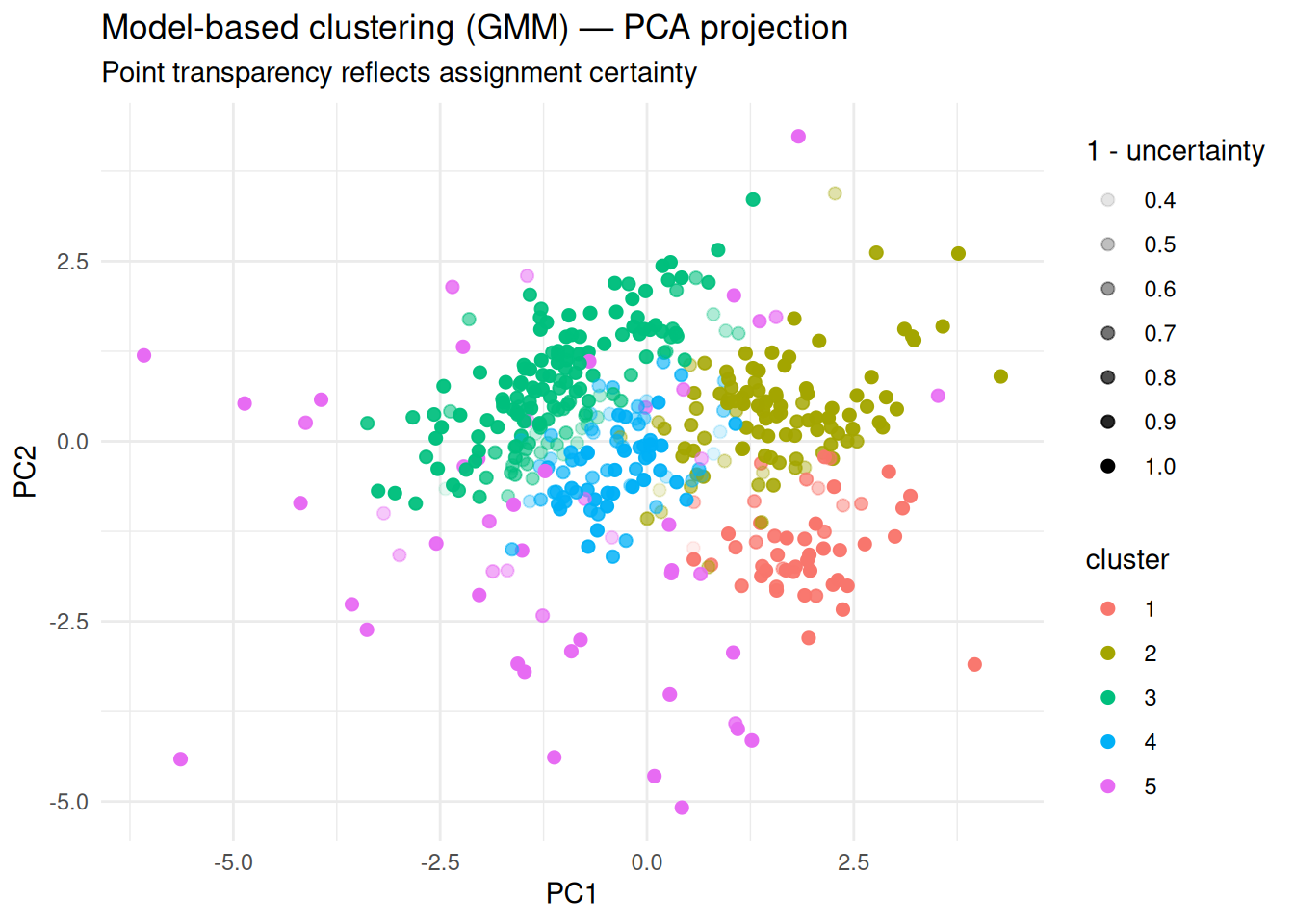
Step 4: Visualization (PCA + uncertainty)
library(ggplot2)
pc <- prcomp(X)
plot_df <- data.frame(
PC1 = pc$x[, 1],
PC2 = pc$x[, 2],
cluster = factor(cl),
uncertainty = uncert
)
ggplot(plot_df, aes(PC1, PC2, color = cluster)) +
geom_point(aes(alpha = 1 - uncertainty), size = 2) +
theme_minimal() +
labs(
title = "Model-based clustering (GMM) — PCA projection",
subtitle = "Point transparency reflects assignment certainty",
x = "PC1", y = "PC2"
)
Step 5: Compare with K-means (same number of clusters)
set.seed(123)
km <- kmeans(X, centers = length(unique(cl)), nstart = 50)
ari <- adjustedRandIndex(cl, km$cluster)
ari## [1] 0.3911639Interpretation:
- If ARI is high, both methods recover similar structure.
- If ARI is lower, investigate covariance shapes: GMM can fit ellipses; K-means cannot.
Step 6: Cluster profiling
df_prof <- as.data.frame(X)
df_prof$cluster <- factor(cl)
profiles <- df_prof |>
group_by(cluster) |>
summarize(across(where(is.numeric), mean), .groups = "drop")
profiles| cluster | Fresh | Milk | Grocery | Frozen | Detergents_Paper | Delicassen |
|---|---|---|---|---|---|---|
| 1 | -1.0655120 | 0.7999948 | 1.0069265 | -1.2232358 | 1.1661900 | -0.2222957 |
| 2 | 0.1712266 | 0.8216657 | 0.8652300 | -0.0062327 | 1.0215707 | 0.4978113 |
| 3 | 0.5462568 | -0.5670315 | -0.5730281 | 0.6567221 | -0.5986691 | 0.1313179 |
| 4 | -0.0650259 | -0.1181184 | -0.2940579 | -0.2959572 | -0.2604843 | -0.1358719 |
| 5 | -0.8145286 | -0.5063557 | -0.5019892 | -0.2847088 | -0.9377190 | -0.9286635 |
For business communication, it is often preferable to profile clusters on original-scale variables (before scaling/log), and report relative differences.
Summary
- Model-based clustering provides a probabilistic framework with soft assignments.
- It naturally supports elliptical clusters and explicit uncertainty quantification.
- BIC-based selection helps choose the number of components.
- Comparing GMM vs K-means clarifies whether “spherical” clustering is too restrictive.
A work by Gianluca Sottile
gianluca.sottile@unipa.it