Ridge & Lasso Regression in R — Penalized Linear Models with glmnet
Ridge & Lasso Regression (Penalized OLS)
Why penalization?
OLS estimates can become unstable when:
- predictors are highly correlated (multicollinearity),
- the number of predictors is large relative to the sample size,
- you want a model that generalizes better (lower variance) and/or performs variable selection.
Penalized regression adds a constraint/penalty to shrink coefficients toward zero, trading a bit of bias for lower variance.
Theory: Ridge, Lasso, Elastic Net
Assume a linear model \(y = X\beta + \epsilon\) with \(X \in \mathbb{R}^{n \times p}\). The OLS objective minimizes RSS: \[ \min_{\beta_0,\beta}\ \frac{1}{2n}\sum_{i=1}^n (y_i - \beta_0 - x_i^T\beta)^2 \]
Ridge (L2 penalty)
\[ \min_{\beta_0,\beta}\ \frac{1}{2n}\sum_{i=1}^n (y_i - \beta_0 - x_i^T\beta)^2\ +\ \lambda \sum_{j=1}^p \beta_j^2 \]
- Shrinks coefficients smoothly.
- Typically does not set coefficients exactly to zero (no hard feature selection).
- Very useful under multicollinearity.
Lasso (L1 penalty)
\[ \min_{\beta_0,\beta}\ \frac{1}{2n}\sum_{i=1}^n (y_i - \beta_0 - x_i^T\beta)^2\ +\ \lambda \sum_{j=1}^p |\beta_j| \]
- Encourages sparsity: some \(\hat\beta_j = 0\).
- Performs variable selection (often easier interpretation).
Elastic Net (mix L1 + L2)
Elastic Net combines both: \[ \min_{\beta_0,\beta}\ \frac{1}{2n}\|y-\beta_0\mathbf{1}-X\beta\|_2^2 \ +\ \lambda\left[(1-\alpha)\frac{1}{2}\|\beta\|_2^2 + \alpha\|\beta\|_1\right] \] with \(0 \le \alpha \le 1\):
- \(\alpha = 0\) → ridge
- \(\alpha = 1\) → lasso
- \(0<\alpha<1\) → elastic net
Hyperparameter intuition
- \(\lambda = 0\): no penalization (≈ OLS).
- \(\lambda \uparrow\): stronger shrinkage; lasso sets more coefficients to zero.
- \(\alpha\): controls sparsity vs stability.
Practice in R with glmnet
Key points:
glmnetexpects a numeric matrixx(usemodel.matrix()for factors).- Use cross-validation to select \(\lambda\) via
cv.glmnet(). - Common choices:
lambda.min: minimizes CV error.lambda.1se: simplest model within 1 standard error (more regularization; often more robust).
Example: Ridge vs Lasso on mtcars (Gaussian regression)
We predict mpg using a mix of numeric and categorical
features.
Step 1: Data preparation (design matrix)
library(dplyr)
df <- mtcars |>
mutate(
cyl = factor(cyl),
am = factor(am, labels = c("auto", "manual")),
vs = factor(vs)
)
# Model matrix: adds intercept column by default; glmnet handles intercept separately,
# so we remove it with "-1" here and let glmnet fit intercept = TRUE.
X <- model.matrix(mpg ~ . - 1, data = df)
y <- df$mpg
dim(X)## [1] 32 12## cyl4 cyl6 cyl8 disp hp drat
## Mazda RX4 0 1 0 160 110 3.90
## Mazda RX4 Wag 0 1 0 160 110 3.90
## Datsun 710 1 0 0 108 93 3.85
## Hornet 4 Drive 0 1 0 258 110 3.08
## Hornet Sportabout 0 0 1 360 175 3.15
## Valiant 0 1 0 225 105 2.76Step 2: Train/test split
Step 3: Baseline OLS (for comparison)
train_ols <- data.frame(mpg = y_train, X_train)
test_ols <- data.frame(mpg = y_test, X_test)
fit_ols <- lm(mpg ~ ., data = train_ols)
summary(fit_ols)##
## Call:
## lm(formula = mpg ~ ., data = train_ols)
##
## Residuals:
## Min 1Q Median 3Q Max
## -3.7677 -1.3928 -0.0578 0.7350 4.6582
##
## Coefficients: (1 not defined because of singularities)
## Estimate Std. Error t value Pr(>|t|)
## (Intercept) 16.57753 19.19814 0.863 0.4048
## cyl4 2.46724 6.70666 0.368 0.7194
## cyl6 -0.20269 4.81932 -0.042 0.9671
## cyl8 NA NA NA NA
## disp 0.02219 0.02786 0.797 0.4411
## hp -0.03423 0.04199 -0.815 0.4308
## drat -0.12025 2.31432 -0.052 0.9594
## wt -4.53231 2.54011 -1.784 0.0997 .
## qsec 0.75808 0.95328 0.795 0.4419
## vs1 1.53834 2.93413 0.524 0.6096
## ammanual 2.47468 2.99684 0.826 0.4251
## gear 0.12976 2.20021 0.059 0.9539
## carb 0.64798 1.23947 0.523 0.6106
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Residual standard error: 3.033 on 12 degrees of freedom
## Multiple R-squared: 0.8818, Adjusted R-squared: 0.7734
## F-statistic: 8.138 on 11 and 12 DF, p-value: 0.0005336pred_ols <- predict(fit_ols, newdata = test_ols)
rmse <- function(y, yhat) sqrt(mean((y - yhat)^2))
rmse_ols <- rmse(y_test, pred_ols)
rmse_ols## [1] 2.1292Step 4: Ridge regression (alpha = 0) with CV
# install.packages("glmnet") # if needed
library(glmnet)
set.seed(123)
cv_ridge <- cv.glmnet(
x = X_train,
y = y_train,
alpha = 0, # ridge
nfolds = 10, # default is 10
standardize = TRUE
)
plot(cv_ridge)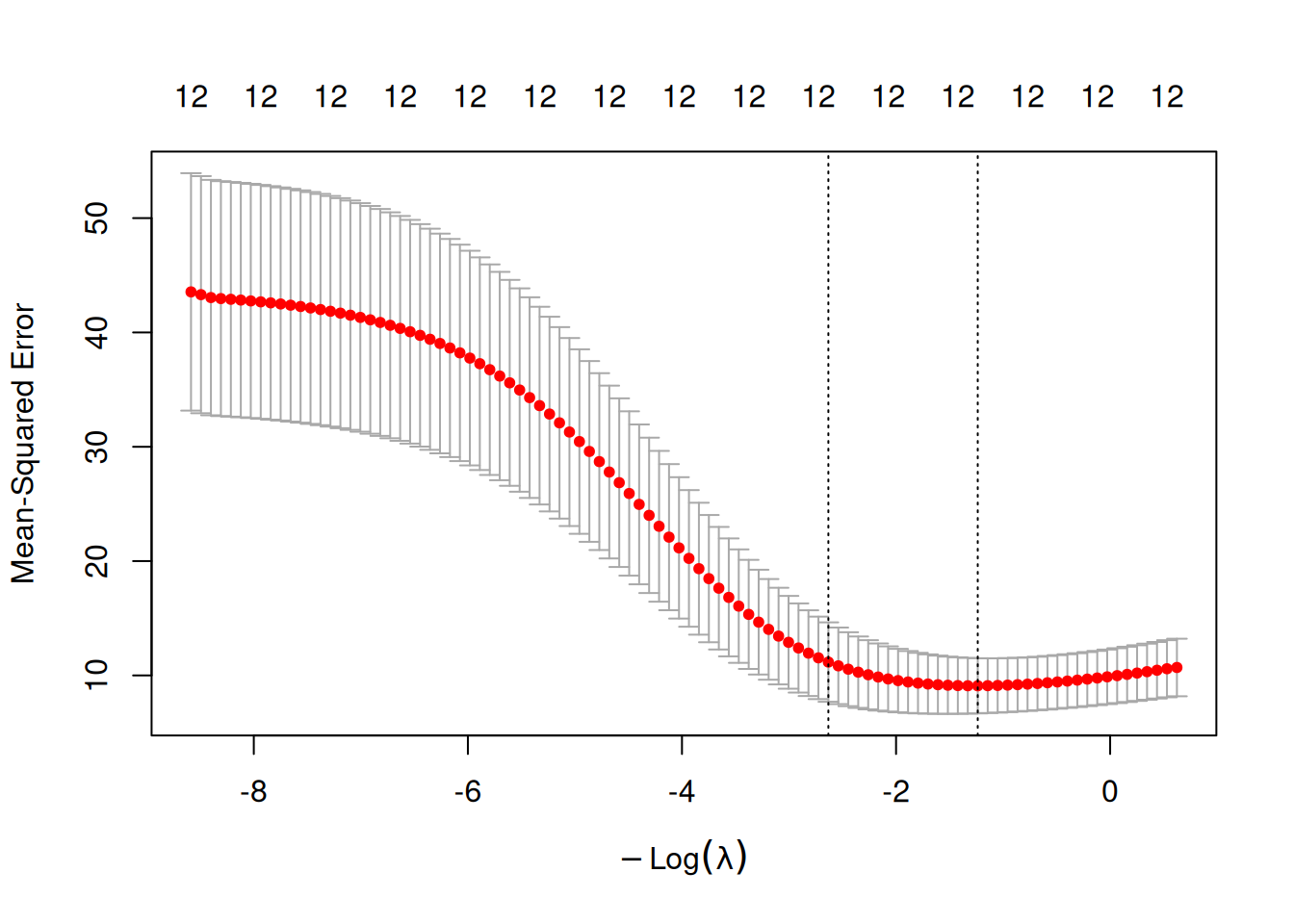
## [1] 3.444694## [1] 13.90629Fit ridge at selected lambda and evaluate:
pred_ridge_min <- predict(cv_ridge, newx = X_test, s = "lambda.min")
pred_ridge_1se <- predict(cv_ridge, newx = X_test, s = "lambda.1se")
rmse_ridge_min <- rmse(y_test, pred_ridge_min)
rmse_ridge_1se <- rmse(y_test, pred_ridge_1se)
c(rmse_ols = rmse_ols, rmse_ridge_min = rmse_ridge_min, rmse_ridge_1se = rmse_ridge_1se)## rmse_ols rmse_ridge_min rmse_ridge_1se
## 2.129200 2.055639 2.218518Inspect coefficients:
## 10 x 1 sparse Matrix of class "dgCMatrix"
## lambda.min
## (Intercept) 20.049804674
## cyl4 1.509365191
## cyl6 -0.715565529
## cyl8 -1.064064671
## disp -0.005891425
## hp -0.010170279
## drat 0.913128419
## wt -1.233903891
## qsec 0.153282689
## vs1 1.011905504Step 5: Lasso regression (alpha = 1) with CV
set.seed(123)
cv_lasso <- cv.glmnet(
x = X_train,
y = y_train,
alpha = 1, # lasso
nfolds = 10,
standardize = TRUE
)
plot(cv_lasso)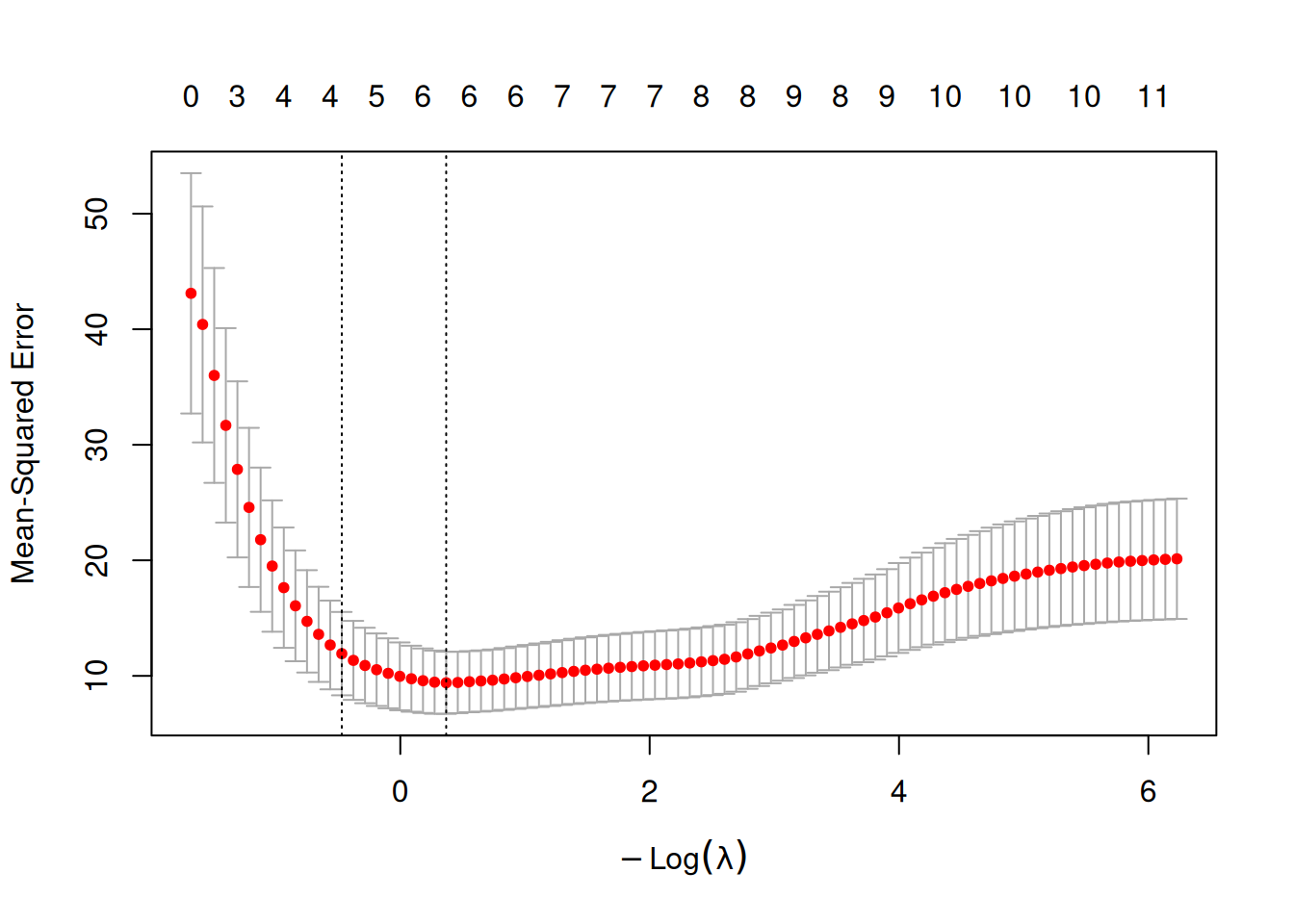
## [1] 0.6921193## [1] 1.598885Evaluate and compare:
pred_lasso_min <- predict(cv_lasso, newx = X_test, s = "lambda.min")
pred_lasso_1se <- predict(cv_lasso, newx = X_test, s = "lambda.1se")
rmse_lasso_min <- rmse(y_test, pred_lasso_min)
rmse_lasso_1se <- rmse(y_test, pred_lasso_1se)
c(
rmse_ols = rmse_ols,
rmse_lasso_min = rmse_lasso_min,
rmse_lasso_1se = rmse_lasso_1se
)## rmse_ols rmse_lasso_min rmse_lasso_1se
## 2.129200 2.060473 2.171117Check sparsity (number of non-zero coefficients):
## [1] 6List selected predictors:
## [1] "(Intercept)" "cyl4" "cyl8" "hp" "wt"
## [6] "vs1"Step 6: Coefficient paths (optional)
fit_lasso_path <- glmnet(X_train, y_train, alpha = 1, standardize = TRUE)
plot(fit_lasso_path, xvar = "lambda", label = TRUE, main = "Lasso coefficient paths")
abline(v = -log(cv_lasso$lambda.min), lty = 2, col = "red")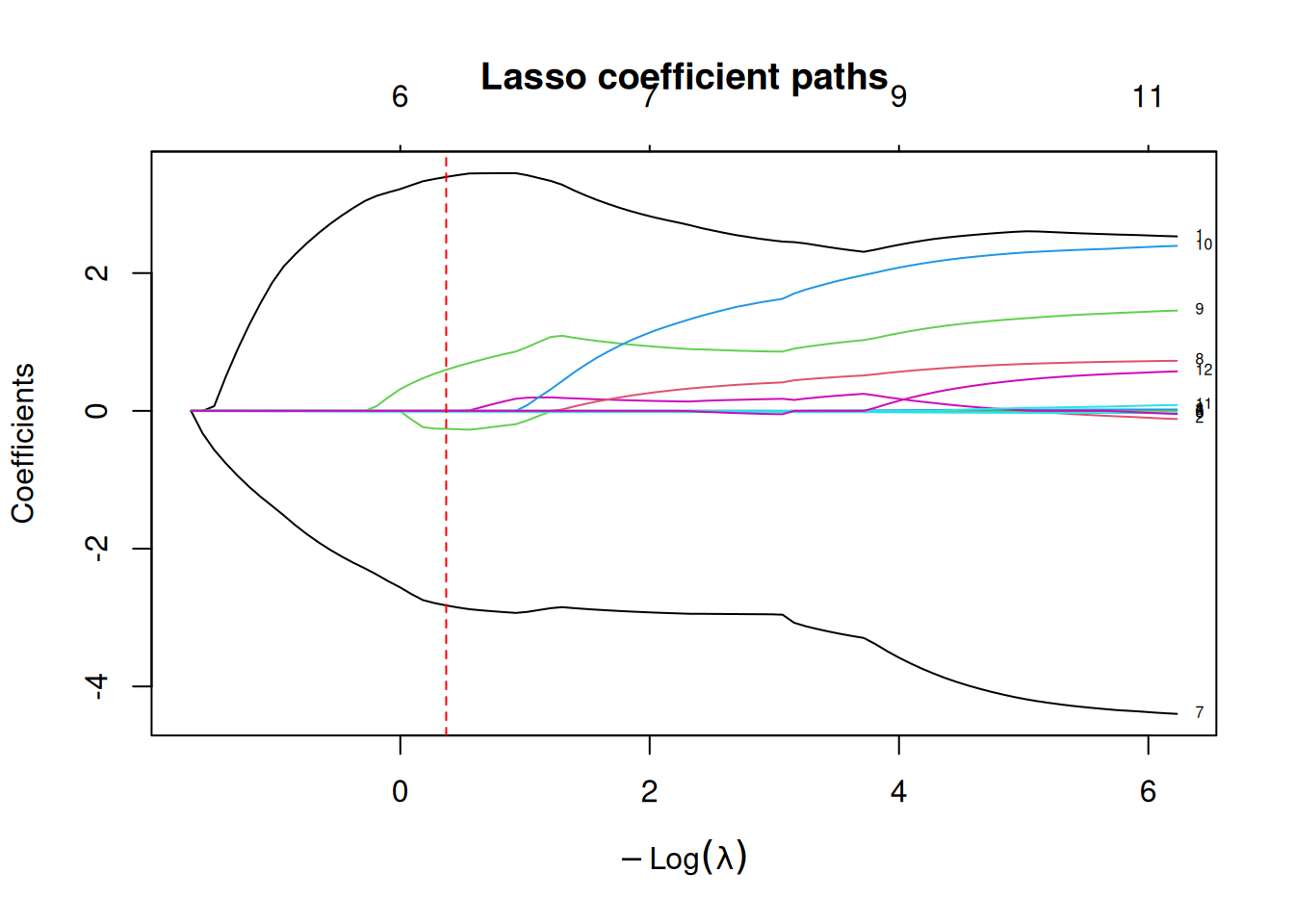
fit_ridge_path <- glmnet(X_train, y_train, alpha = 0, standardize = TRUE)
plot(fit_ridge_path, xvar = "lambda", label = FALSE, main = "Ridge coefficient paths")
abline(v = -log(cv_ridge$lambda.min), lty = 2, col = "red")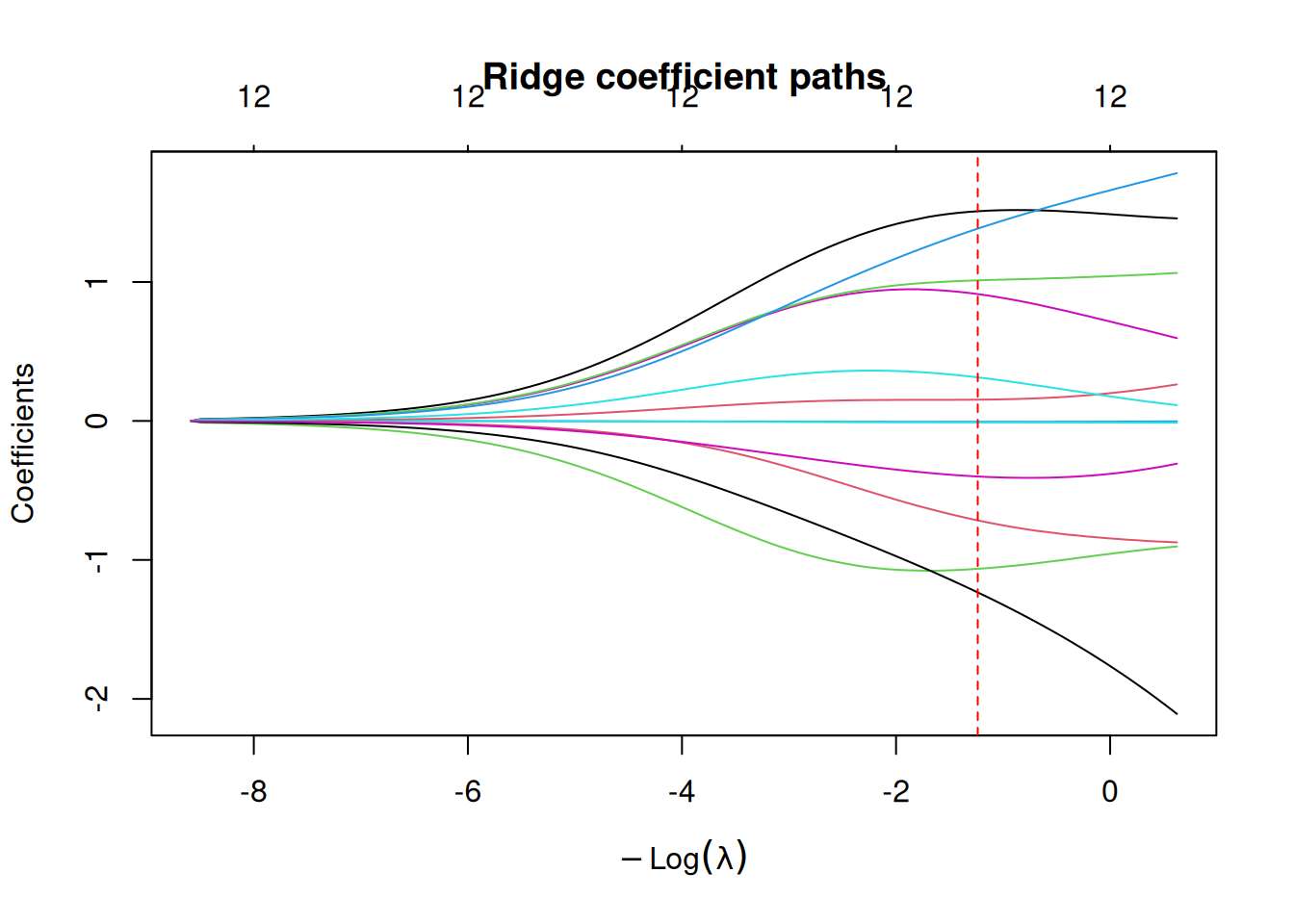
Optional: Elastic Net (alpha between 0 and 1)
If predictors are highly correlated, elastic net can outperform pure lasso.
set.seed(123)
cv_enet <- cv.glmnet(
x = X_train,
y = y_train,
alpha = 0.5, # elastic net
nfolds = 10,
standardize = TRUE
)
plot(cv_enet)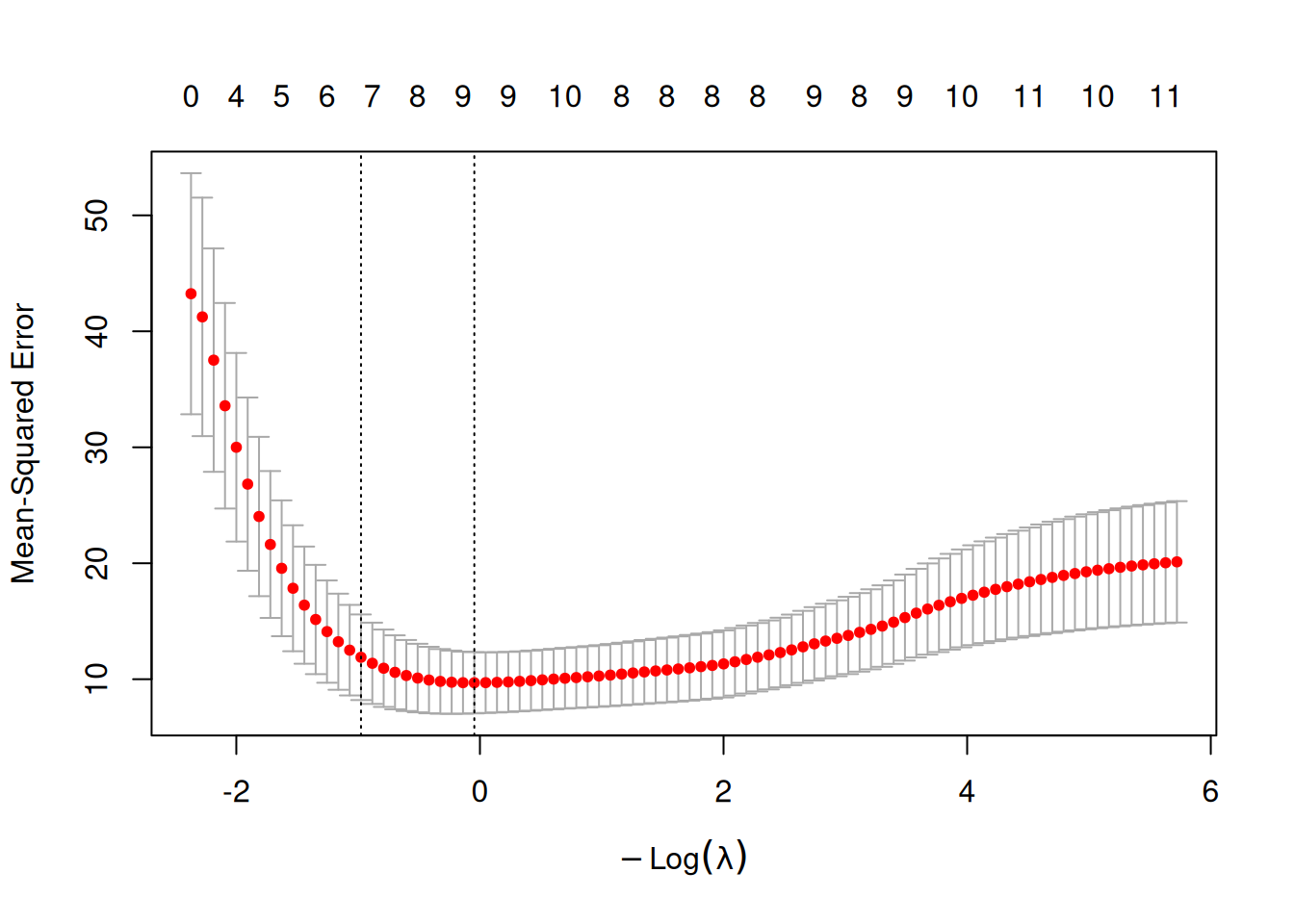
pred_enet <- predict(cv_enet, newx = X_test, s = "lambda.min")
rmse_enet <- rmse(y_test, pred_enet)
c(rmse_lasso_min = rmse_lasso_min, rmse_ridge_min = rmse_ridge_min, rmse_enet_min = rmse_enet)## rmse_lasso_min rmse_ridge_min rmse_enet_min
## 2.060473 2.055639 2.077020Best practices & warnings
- Always use cross-validation for \(\lambda\); prefer
lambda.minwhen you want stability and parsimony. - Standardization is important so that the penalty treats predictors comparably.
- Lasso can be unstable when predictors are strongly correlated (it may pick one arbitrarily); elastic net is often safer.
- If the goal is inference (p-values, CI), penalized regression needs extra care (post-selection inference); treat as predictive by default.
Summary
You learned how to: - Define ridge/lasso/elastic net with penalized
objectives. - Fit ridge and lasso in R using glmnet +
cv.glmnet(). - Select \(\lambda\) via CV (lambda.min
vs lambda.1se). - Compare test RMSE and interpret shrinkage
vs sparsity.
Penalized regression is a modern alternative to stepwise selection for regression problems with many correlated predictors.
A work by Gianluca Sottile
gianluca.sottile@unipa.it