Spectral Clustering in R — When to Switch Families (Graph-based Clustering)
Spectral Clustering
Spectral clustering is a graph-based method. It builds an affinity graph \(W\) from similarities (e.g., RBF kernel): \(W_{ij} = \exp(-\|x_i - x_j\|^2 / (2\sigma^2))\).
Define the degree matrix \(D\) where \(D_{ii} = \sum_j W_{ij}\), and (one version of) the normalized Laplacian: \(L = I - D^{-1/2} W D^{-1/2}\).
The algorithm embeds data into the space spanned by a few eigenvectors of \(L\) (or a related matrix) and then runs a simple clustering method (often K-means) in that embedding.
This family is particularly useful when clusters are not well-separated in Euclidean space but are separable by connectivity on a similarity graph.
In this lesson:
- Fit spectral clustering with an RBF kernel.
- Tune the kernel scale \(\sigma\) pragmatically.
- Compare with K-means/hierarchical results.
Step 1: Import + preprocessing
library(readr)
library(dplyr)
local_path <- "raw_data/wholesale_customers.csv"
df_raw <- read_csv(local_path, show_col_types = FALSE)
spend_vars <- c("Fresh", "Milk", "Grocery", "Frozen", "Detergents_Paper", "Delicassen")
X <- df_raw |>
mutate(across(all_of(spend_vars), ~ log1p(.x))) |>
select(all_of(spend_vars)) |>
mutate(across(everything(), scale)) |>
as.matrix()Step 2: Choose k (reuse from prior analyses or evaluate a grid)
Spectral clustering typically requires specifying \(k\). Here we illustrate a simple approach: test a small grid and select based on silhouette in the embedded labels.
library(cluster)
library(kernlab)
avg_sil_from_labels <- function(X, labels) {
if (length(unique(labels)) < 2) return(NA_real_)
s <- silhouette(labels, dist(X))
mean(s[, 3])
}
k_grid <- 2:10
sigma_grid <- c(0.1, 0.2, 0.5, 1)
results <- expand.grid(k = k_grid, sigma = sigma_grid)
results$avg_sil <- NA_real_
set.seed(123)
for (i in seq_len(nrow(results))) {
k <- results$k[i]
sig <- results$sigma[i]
sc <- specc(X, centers = k, kernel = "rbfdot", kpar = list(sigma = sig))
results$avg_sil[i] <- avg_sil_from_labels(X, as.integer(sc))
}
results <- results[order(-results$avg_sil), ]
head(results, 10)| k | sigma | avg_sil | |
|---|---|---|---|
| 28 | 2 | 1.0 | 0.2897449 |
| 10 | 2 | 0.2 | 0.2892797 |
| 19 | 2 | 0.5 | 0.2888352 |
| 1 | 2 | 0.1 | 0.2868777 |
| 11 | 3 | 0.2 | 0.2498515 |
| 2 | 3 | 0.1 | 0.2071218 |
| 12 | 4 | 0.2 | 0.1920744 |
| 21 | 4 | 0.5 | 0.1790377 |
| 3 | 4 | 0.1 | 0.1777185 |
| 5 | 6 | 0.1 | 0.1729661 |
Select the best configuration (highest silhouette), and then refit:
best_k <- results$k[1]
best_sigma <- results$sigma[1]
set.seed(123)
sc_best <- specc(X, centers = best_k, kernel = "rbfdot", kpar = list(sigma = best_sigma))
cl_sc <- as.integer(sc_best)
table(cl_sc)## cl_sc
## 1 2
## 254 186Step 3: Visualization (PCA projection)
library(ggplot2)
pc <- prcomp(X)
plot_df <- data.frame(
PC1 = pc$x[, 1],
PC2 = pc$x[, 2],
cluster = factor(cl_sc)
)
ggplot(plot_df, aes(PC1, PC2, color = cluster)) +
geom_point(size = 2, alpha = 0.85) +
theme_minimal() +
labs(
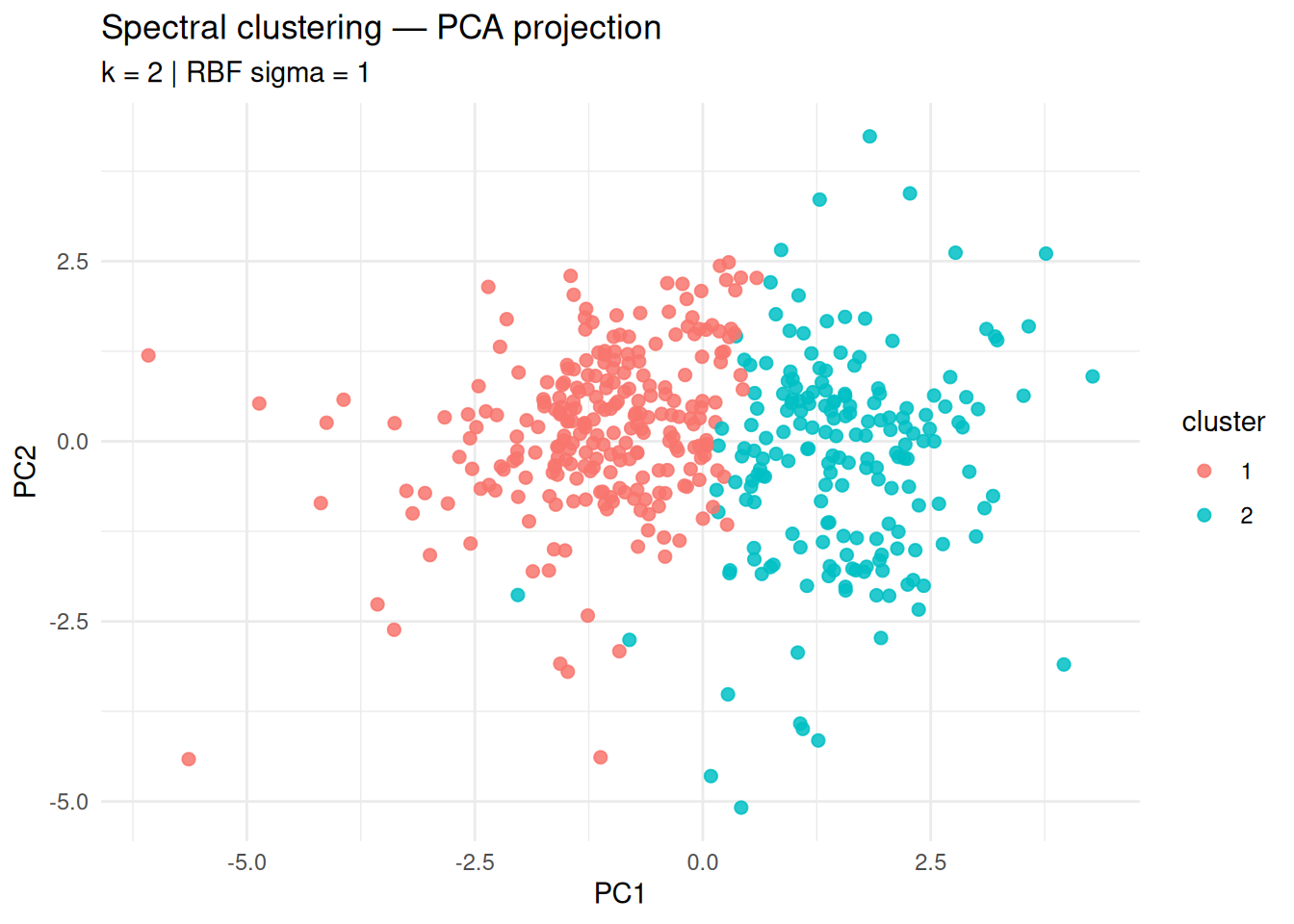
title = "Spectral clustering — PCA projection",
subtitle = paste("k =", best_k, "| RBF sigma =", best_sigma),
x = "PC1", y = "PC2"
)
Step 4: Comparison with K-means (same k)
library(mclust)
set.seed(123)
km <- kmeans(X, centers = best_k, nstart = 50)
ari <- adjustedRandIndex(cl_sc, km$cluster)
ari## [1] 0.893599Interpretation:
- Spectral clustering is preferred when K-means struggles due to non-convex or graph-structured clusters.
- If ARI is high, it suggests the structure is already well captured by centroid-based clustering.
Step 5: Cluster profiles
df_prof <- as.data.frame(X)
df_prof$cluster <- factor(cl_sc)
profiles <- df_prof |>
group_by(cluster) |>
summarize(across(where(is.numeric), mean), .groups = "drop")
profiles| cluster | Fresh | Milk | Grocery | Frozen | Detergents_Paper | Delicassen |
|---|---|---|---|---|---|---|
| 1 | 0.2152193 | -0.5720568 | -0.6377129 | 0.2678828 | -0.6509132 | -0.1329263 |
| 2 | -0.2939016 | 0.7811958 | 0.8708552 | -0.3658184 | 0.8888815 | 0.1815230 |
Summary
- Spectral clustering works on a similarity graph and can capture structures that are not well modeled by spherical clusters.
- The kernel scale \(\sigma\) is crucial; a small grid search with an internal index (e.g., silhouette) is a practical approach.
- Compare against K-means to justify switching clustering families rather than forcing centroid-based methods.
A work by Gianluca Sottile
gianluca.sottile@unipa.it