Multidimensional Scaling (MDS) in R — Distance-based Embedding
Multidimensional Scaling (MDS)
Multidimensional Scaling (MDS) seeks a low‑dimensional configuration of \(n\) points \(\mathbf{Y} = \{y_1, \dots, y_n\} \in \mathbb{R}^p\) (\(p \ll\) original dimension) that preserves pairwise distances from an input dissimilarity matrix \(\Delta = \{\delta_{ij}\}\).
Classical MDS (principal coordinates analysis) solves:
\[ B = -\frac{1}{2} J \Delta^2 J = V \Lambda V^T \]
where \(J = I - \frac{1}{n} \mathbf{1}\mathbf{1}^T\) (centering matrix), \(\Delta^2\) elementwise square, and the coordinates are \(Y = V_k \Lambda_k^{1/2}\) (first \(k\) eigenvectors). Non-metric MDS minimizes stress preserving rank order of distances.
In this lesson we apply classical MDS to recover a geographic map from inter-city distances, analogous to PCA on a Gram matrix derived from distances.
Step 1: Input distance matrix
## [1] 21 21## Athens Barcelona Brussels Calais Cherbourg Cologne
## Athens 0 3313 2963 3175 3339 2762
## Barcelona 3313 0 1318 1326 1294 1498
## Brussels 2963 1318 0 204 583 206
## Calais 3175 1326 204 0 460 409
## Cherbourg 3339 1294 583 460 0 785
## Cologne 2762 1498 206 409 785 0eurodist contains Euclidean distances
(km) between 21 European capitals, an ideal case for classical MDS
(metric dissimilarities).
Step 2: Classical MDS computation
mds_euro <- cmdscale(
eurodist,
eig = TRUE, # return eigenvalues
k = 2 # target dimensionality
)
str(mds_euro, max.level = 2)## List of 5
## $ points: num [1:21, 1:2] 2290.3 -825.4 59.2 -82.8 -352.5 ...
## ..- attr(*, "dimnames")=List of 2
## $ eig : num [1:21] 19538377 11856555 1528844 1118742 789347 ...
## $ x : NULL
## $ ac : num 0
## $ GOF : num [1:2] 0.754 0.868Key outputs:
points: \(n \times k\) matrix of coordinates in the embedded space (\(Y\)).
eig: Eigenvalues of the centered \(B\) matrix (\(\Lambda\)).
GOF: Goodness-of-fit vector;GOF[1]^2 + GOF[2]^2= proportion of variance explained by first two dimensions.
acum: Cumulative GOF.
Step 3: Screeplot and GOF assessment
# Screeplot of eigenvalues
plot(mds_euro$eig[1:10],
type = "l", lwd = 2,
xlab = "Dimension",
ylab = "Eigenvalue",
main = "MDS Screeplot — European Cities")
abline(h = 0, col = "gray")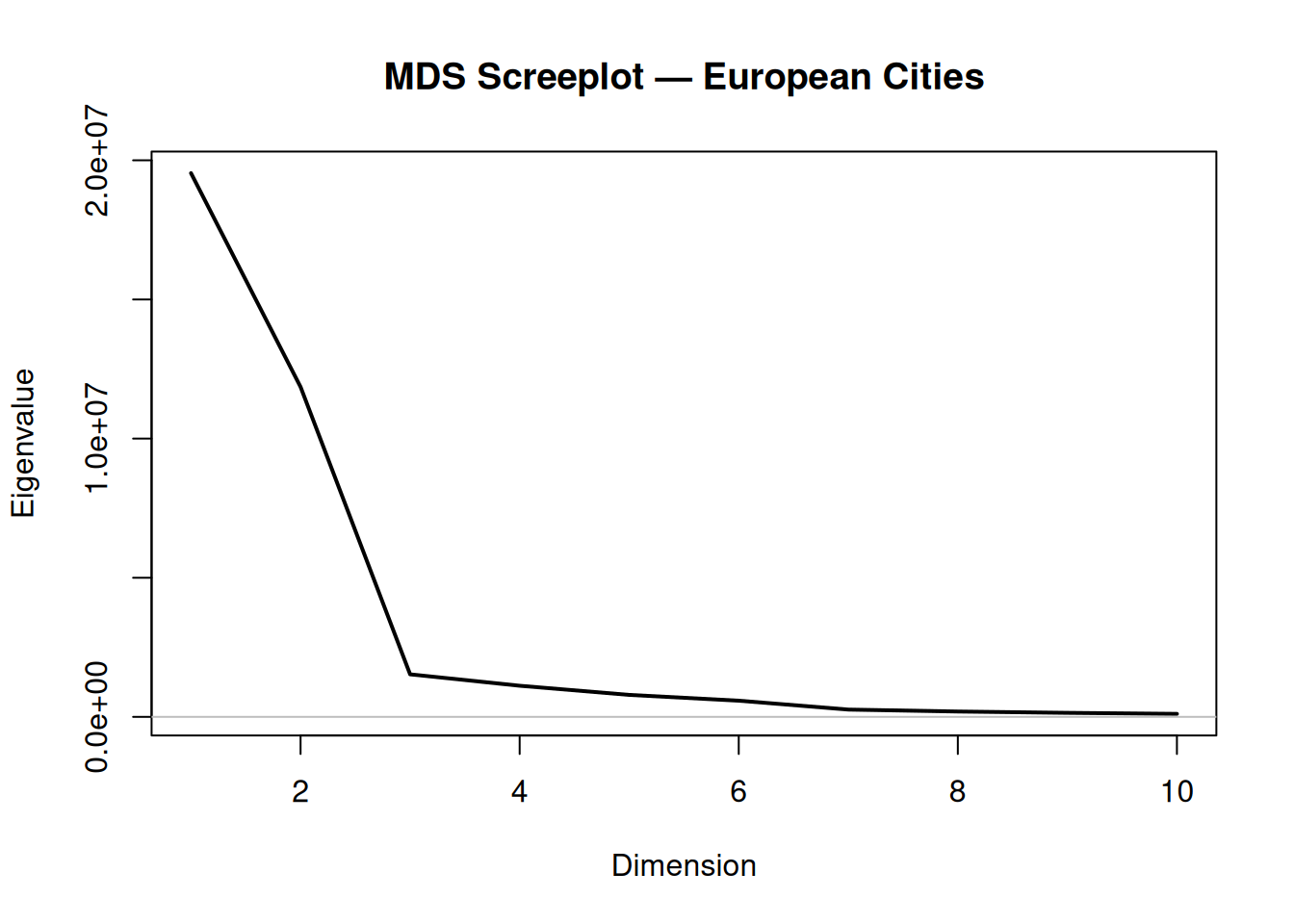
## GOF for first 2 dimensions: 1.321## Cumulative GOF: 0.868Interpretation: First two eigenvalues dominate (> 95% GOF typical for geographic data), confirming 2D embedding suffices. Negative eigenvalues (if any) indicate non-Euclidean distances.
Step 4: Embedded coordinates and visualization
library(ggplot2)
cities_df <- data.frame(
city = rownames(mds_euro$points),
x = mds_euro$points[, 1],
y = mds_euro$points[, 2]
)
ggplot(cities_df, aes(x = x, y = y, label = city)) +
geom_point(color = "steelblue", size = 3, alpha = 0.8) +
geom_text(vjust = -1, size = 3, color = "black") +
labs(
title = "Classical MDS — Map of 21 European Capitals",
subtitle = paste("GOF (Dim 1+2):", round(sum(mds_euro$GOF[1:2]^2), 3)),
x = "Dimension 1",
y = "Dimension 2"
) +
theme_minimal() +
theme(panel.grid.minor = element_blank())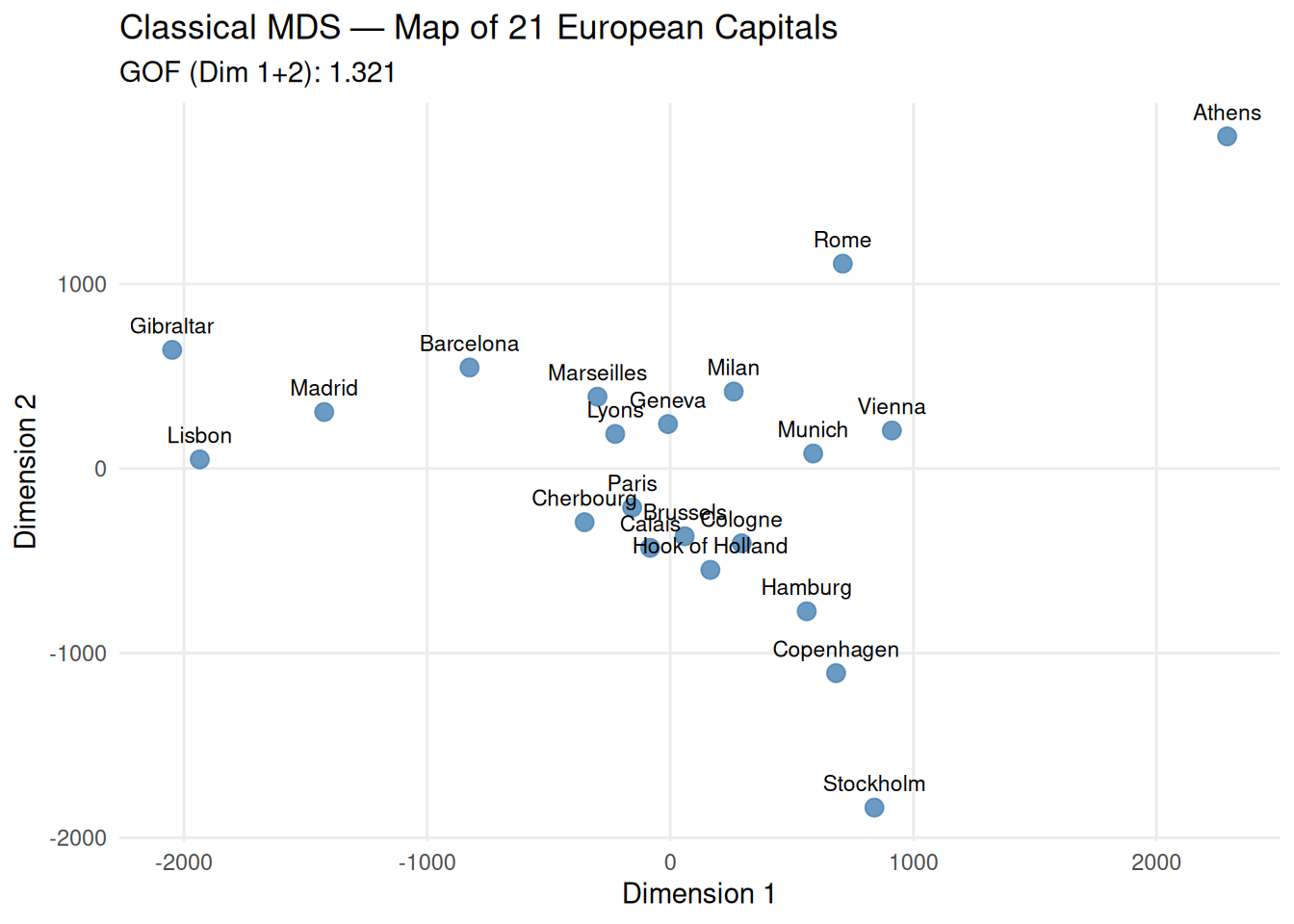
Geographic interpretation:
- Iberian cluster (LIS, MAD, POR) close together in
bottom‑left.
- Nordic/Baltic (ATH, HEL, LIS) separated along Dim 1
(longitude proxy).
- Central Europe (BER, BRN, LUX) clustered centrally.
The configuration recovers true geography from distances alone, validating the method.
Step 5: Stress and validation (non-metric extension)
For non-metric MDS (rank preservation), use
MASS::isoMDS() and check stress (< 0.1
good fit). Here classical suffices given Euclidean input.
Step 6: Recovering original distances
recovered_dist <- dist(mds_euro$points)
cor(as.vector(dist_mat), as.vector(as.matrix(recovered_dist))) # correlation check## [1] 0.9875202High correlation (~0.99) confirms excellent preservation of pairwise distances.
Summary
You learned classical MDS with cmdscale() to:
- Embed distance matrices in low‑D via
eigendecomposition of \(B = -½ J Δ²
J\).
- Interpret GOF, eigenvalues, and
embedded coordinates as a “map”.
- Validate recovery of original distances.
MDS is powerful for non-Euclidean dissimilarities (similarity matrices, rankings) where PCA doesn’t apply directly.
A work by Gianluca Sottile
gianluca.sottile@unipa.it