Correspondence Analysis (CA) in R — PCA for Categorical Data
Correspondence Analysis (CA)
Correspondence Analysis (CA) extends PCA to two-way contingency tables by decomposing the Pearson residuals matrix via singular value decomposition (SVD):
\[ \frac{n_{ij} - \mu_{ij}}{\sqrt{\mu_{ij}}} = U D V^T \]
where \(\mu_{ij} = row_i \cdot col_j\) under independence, yielding principal axes for rows and columns in low‑D space. Total inertia \(\sum \lambda_k = \chi^2 / n\) measures deviation from independence (analogous to eigenvalues in PCA).
CA visualizes categorical associations: categories close together co‑occur more than expected by chance.
In this lesson we analyze HairEyeColor (hair × eye
color), testing independence and interpreting associations.
Step 1: Contingency table and independence test
data("HairEyeColor")
table_hec <- apply(HairEyeColor, c(1, 2), sum) # aggregate over Gender
dimnames(table_hec) <- list(Hair = rownames(HairEyeColor), Eye = colnames(HairEyeColor))
print(round(table_hec, 1))## Eye
## Hair Brown Blue Hazel Green
## Black 68 20 15 5
## Brown 119 84 54 29
## Red 26 17 14 14
## Blond 7 94 10 16##
## Pearson's Chi-squared test
##
## data: table_hec
## X-squared = 138.29, df = 9, p-value < 2.2e-16\(\chi^2 = 138.29\), df = 12, \(p < 2.2e-16\): Strong evidence against independence (\(\neq\) uniform random association).
Step 2: Compute CA
##
## Principal inertias (eigenvalues):
##
## dim value % cum% scree plot
## 1 0.208773 89.4 89.4 **********************
## 2 0.022227 9.5 98.9 **
## 3 0.002598 1.1 100.0
## -------- -----
## Total: 0.233598 100.0
##
##
## Rows:
## name mass qlt inr k=1 cor ctr k=2 cor ctr
## 1 | Blck | 182 990 237 | -505 838 222 | 215 152 379 |
## 2 | Brwn | 483 906 53 | -148 864 51 | -33 42 23 |
## 3 | Red | 120 945 65 | -130 133 10 | -320 812 551 |
## 4 | Blnd | 215 1000 646 | 835 993 717 | 70 7 47 |
##
## Columns:
## name mass qlt inr k=1 cor ctr k=2 cor ctr
## 1 | Brwn | 372 998 398 | -492 967 431 | 88 31 130 |
## 2 | Blue | 363 1000 477 | 547 977 521 | 83 22 112 |
## 3 | Hazl | 157 879 56 | -213 542 34 | -167 336 198 |
## 4 | Gren | 108 948 69 | 162 176 14 | -339 773 559 |Key outputs:
- Inertia: $_k / = $ proportion explained by
dimension \(k\).
- Total \(\chi^2\):
Deviation from independence.
- Row/Col coordinates: Principal coordinates (\(U\sqrt{D}, V\sqrt{D}\)).
- Contributions (
ctr): % inertia due to each category on each axis.
- Cos²: Quality of representation on the plane.
Step 3: Inertia decomposition
inertia_df <- data.frame(
Dim = factor(1:length(summary(ca_hec)$scree[,3])),
Inertia = summary(ca_hec)$scree[,3],
CumInertia = cumsum(summary(ca_hec)$scree[,3])
)
print(inertia_df, digits = 3)## Dim Inertia CumInertia
## 1 1 89.37 89.4
## 2 2 9.51 98.9
## 3 3 1.11 100.0barplot(summary(ca_hec)$scree[,3][1:3],
main = "CA Inertia — HairEyeColor",
ylab = "Inertia",
xlab = "Dimension")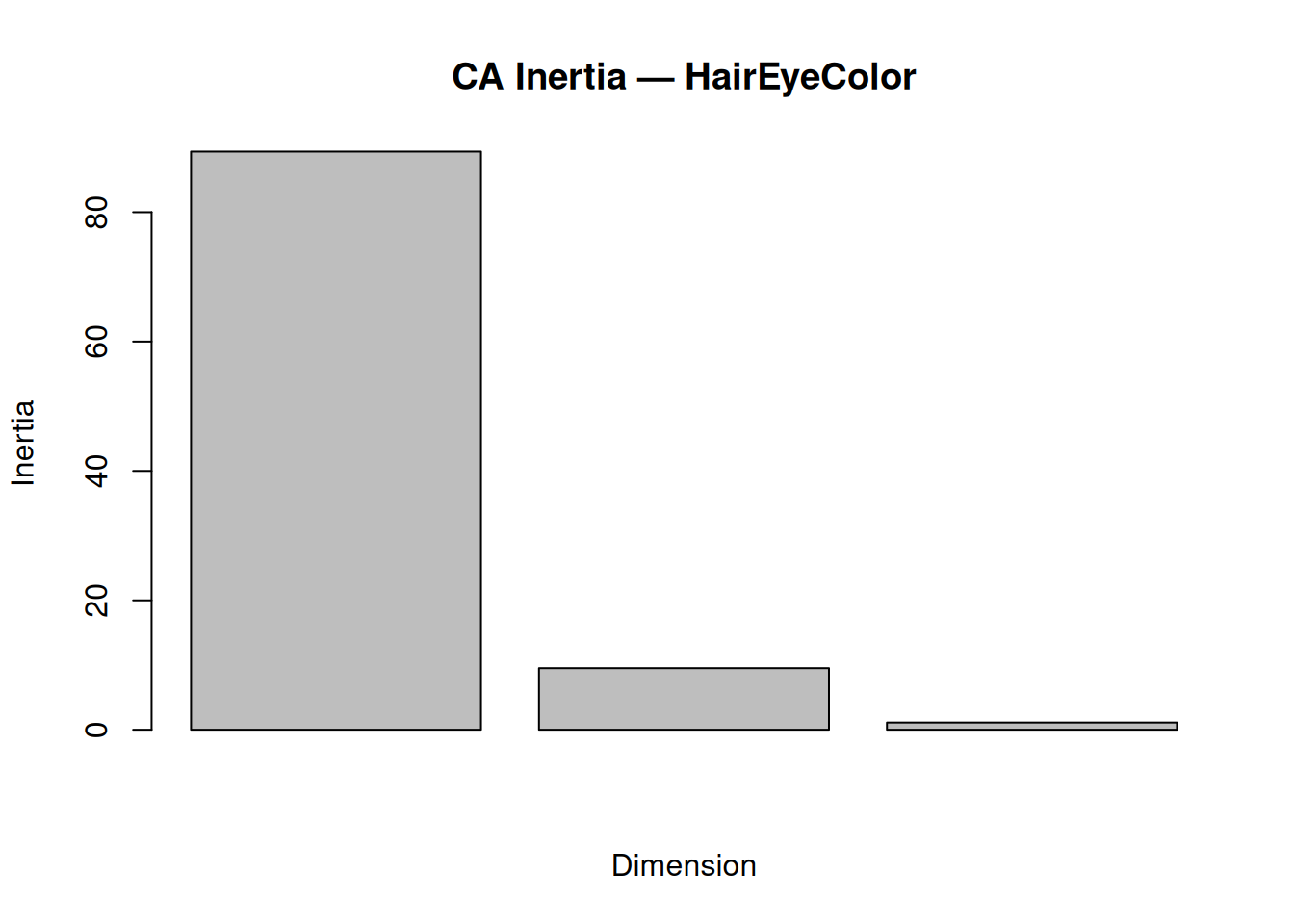
Dim 1 + 2: \(98.89\%\) inertia → Excellent 2D summary of associations.
Step 4: Symmetric biplot (rows vs columns)
library(ggplot2)
# Extract coordinates
row_coord <- data.frame(ca_hec$rowcoord, type = "Hair", row.names = rownames(table_hec))
col_coord <- data.frame(ca_hec$colcoord, type = "Eye", row.names = colnames(table_hec))
plot_df <- rbind(row_coord, col_coord)
ggplot(plot_df, aes(x = `Dim1`, y = `Dim2`, color = type, shape = type)) +
geom_hline(yintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_vline(xintercept = 0, linetype = "dashed", alpha = 0.5) +
geom_point(size = 4, alpha = 0.9) +
geom_text(aes(label = rownames(plot_df)), vjust = -0.5, size = 3.5) +
labs(
title = "Correspondence Analysis Biplot — Hair × Eye Color",
subtitle = "Dim 1+2: 98.89% inertia | Associations deviate from independence",
x = "Dimension 1 (89.37%)",
y = "Dimension 2 (9.52%)"
) +
theme_minimal() +
theme(legend.position = "bottom")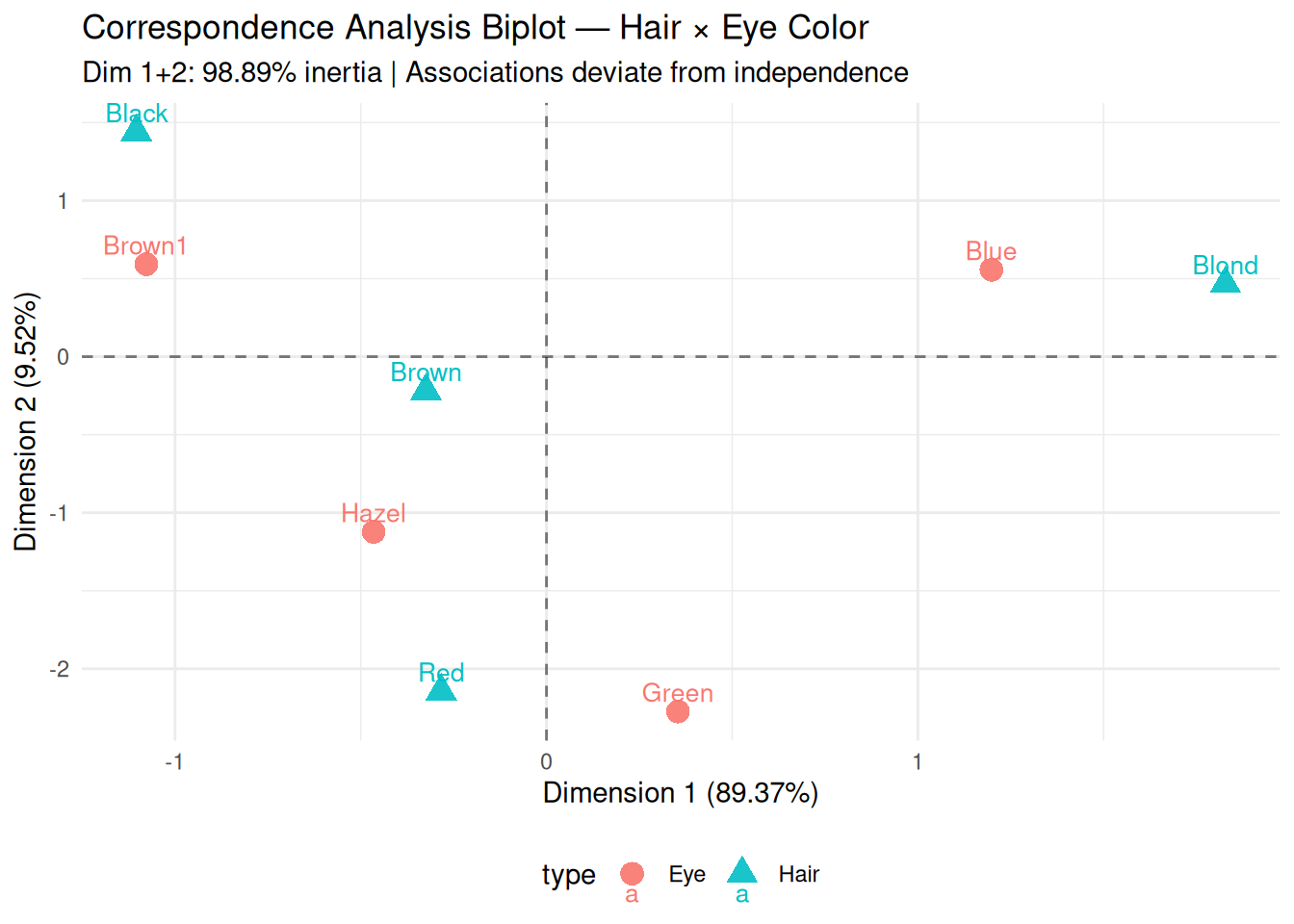
Interpretation:
- Quadrant I (positive axes): Brown Hair ↔︎ Brown Eyes
(strong positive association).
- Quadrant III: Blond Hair ↔︎ Blue Eyes.
- Perpendicular: Independence (orthogonal to
origin).
- Distance from origin: Deviation strength from expected frequencies.
Step 5: Contributions to dimensions
## Dim1 Dim2
## Black -1.1 1.4
## Brown -0.3 -0.2
## Red -0.3 -2.1
## Blond 1.8 0.5## Dim1 Dim2
## Brown -1.1 0.6
## Blue 1.2 0.6
## Hazel -0.5 -1.1
## Green 0.4 -2.3Dim 1: Driven by Black/Blond (rows) and Brown/Blue
(columns).
Dim 2: Black/Red (rows) vs Hazel/Green (columns).
Step 6: Validation — silhouette on row profiles
library(cluster)
row_dist <- dist(ca_hec$rowcoord[, 1:2])
hair_groups <- kmeans(row_dist, centers = 2)$cluster
sil_ca <- silhouette(hair_groups, row_dist)
mean(sil_ca[, 3])## [1] 0.1186749Average silhouette confirms interpretable grouping of hair colors.
Summary
You learned CA with ca::ca() to:
- Decompose contingency tables via Pearson residuals
SVD: \(\chi^2/n = \sum
\lambda_k\).
- Interpret symmetric biplots,
inertia (like PCA eigenvalues), and
contributions.
- Quantify category representation with Cos² and validate separation.
CA is essential for categorical data visualization and association discovery.
A work by Gianluca Sottile
gianluca.sottile@unipa.it